DNA Transcription
In the organisms (viz., prokaryotes and eukaryotes), where coded genetic information is contained in the DNA molecule, different genetically controlled functions of their cells are performed by a different kind of nucleic acid, called non-genetic ribonucleic acid (RNA).
- In the cells of such organisms, the DNA molecule occurs in the chromosomes of the nucleus or nucleoids of eukaryotes or prokaryotes respectively, and the process of protein synthesis occurs in the cytoplasm.
- It is investigated that a DNA molecule does not leave the nucleus to participate directly in the process of protein synthesis but employs different types of non-genetic ribonucleic acid molecules for carrying its genetic information from the nucleus to the site of protein synthesis, i.e., ribosomes.
- Thus, gene expression is accomplished by the transfer of genetic information from DNA to RNA molecules and then from RNA to protein molecules.
- RNA molecules are synthesized by using the base sequence (triplet codons) of one strand of DNA as a template in a polymerization reaction that is catalyzed by enzymes called DNA-dependent RNA polymerases or simply RNA polymerases.
- The process by which RNA molecules are initiated, elongated, and terminated is called transcription.
Chemical Composition Of Non-Genetic Ribonucleic Acid (RNA)
Chemically, the non-genetic RNA is closely related to DNA. However, the non-genetic RNA is a single-stranded polymer of ribonucleotide units.
- It is composed of phosphoric acid, ribose (pentose) sugar, and nitrogen bases, which are purines (adenine and guanine) and pyrimidines (cytosine and uracil).
- The molecular weights of the various RNA molecules vary from about 25,000 to over one million. Its base composition does not follow the base composition of DNA. although, in general, the proportion of A+C roughly equals G+U.
Base Ratios Of Rna From Various Sources (As Molar Percentages):
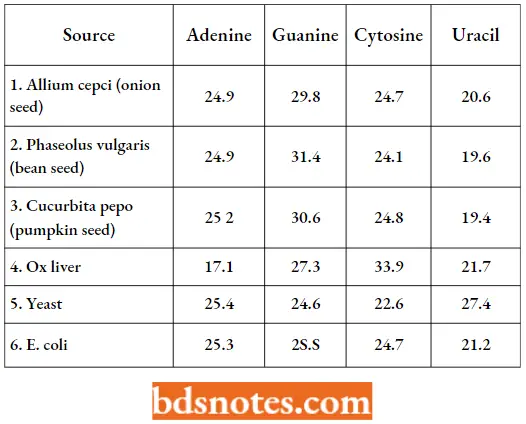
Further, some RNA molecules contain significant proportions of some methylated and an unusual nucleoside known as pseudouridine, in which the glycosidic bond is associated with position 5 of uracil rather than position 3.
“Understanding RNA polymerase through FAQs: Q&A explained”
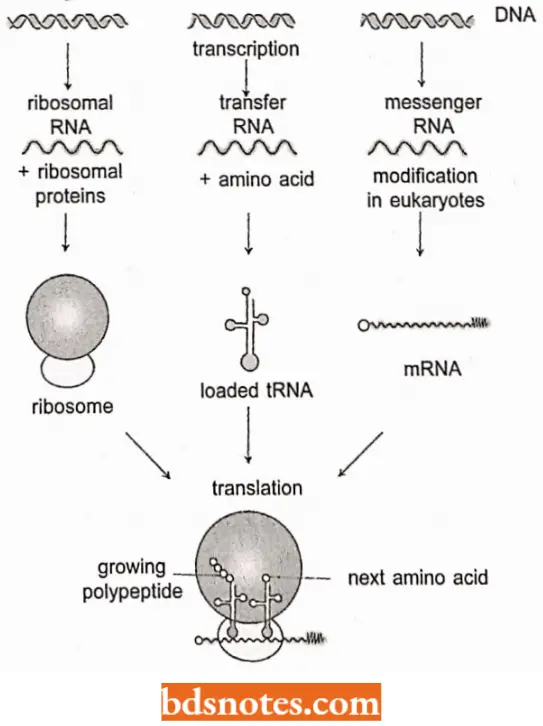
Types of RNA: In a cell, there are three different kinds of RNA which serve three different roles in the process of protein synthesis.
- The first type is messenger RNA (mRNA) which carries the DNA sequence information to particles in the cytoplasm known as ribosomes, where the messenger RNA is translated.
- The second type is transfer RNA (tRNA), which brings the amino acids to the ribosomes, where protein synthesis takes place. The third type of RNA is a structural and functional part of the ribosome called ribosomal RNA (rRNA).
- The general relationship of the roles of these three types of RNA is diagrammed. In addition, small RNAs play other roles in cellular metabolism.
- Further, DNA does not take part directly in protein synthesis because, in eukaryotes, translation occurs in the cytoplasm, whereas DNA remains in the nucleus.
- Molecular biologists suspected for a long time that the genetic intermediate in bacteria and eukaryotes is RNA because the cytoplasmic RNA concentration increases with increasing protein synthesis, and the cytoplasmic RNAs carry nucleotide sequences complementary to the cells DNA.
- Proof of an RNA intermediate came when it was shown that mRNA directs protein synthesis.
Comparison Between DNA Replication And Transcription
Replication and transcription are two chief activities of DNA and they should be compared at the outset in the following manner:
- In DNA replication an enzyme (or enzyme complex) matches up complementary deoxyribonucleoside triphosphates with a DNA template according to Watson-Crick pairing rules and the bases are polymerized to form a daughter DNA molecule.
- In DNA transcription a different enzyme, called DNA-dependent RNA polymerase medicates a similar process, except that complementary ribonucleoside-triphosphates are matched with the DNA template, and RNA polymers are formed.
- The base uracil (U) replaces thymine in RNA but, otherwise, the DNA strand is faithfully copied. The RNA copy has the opposite polarity from the template, so that, for example, the sequence
 in DNA is
in DNA is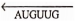 transcribes as in RNA.
transcribes as in RNA. - The RNA transcripts do not ordinarily remain hydrogen-bonded with their DNA templates as DNA daughter strands do. Instead, they “peel off’ the template as they are formed, thus, becoming available to participate in protein synthesis.
- DNA of prokaryotes (E.coli) and eukaryotes have one or few initiation points for DNA replication, so that, at least, large portions of the genome are copied into single, enormous daughter DNA molecules.
- In contrast, the synthesis of RNA molecules is initiated at close intervals along the DNA and, thus, relatively short RNA copies are produced.
- Lastly, replication and transcription have to serve two different functions.
- The purpose of replication is to conserve the entire genome for the next generation, whereas the purpose of transcription is to make RNA copies of individual genes that cells can use in the metabolism.
- The copies are themselves not endowed with the same kind of permanence as the genetic material; instead, they are typically degraded by cellular nucleases, once their functional usefulness has been spent.
Prokaryotic Transcription
Transcription Involves The Following Three Aspects:
- The enzymatic synthesis of RNA;
- The signals that determine at what points on a DNA molecule transcription starts and stops; and
- The types of transcription products and how they are converted to the RNA molecules needed by the cell.
Enzymatic Synthesis Of RNA: The essential chemical characteristics of the synthesis of RNA are the following:
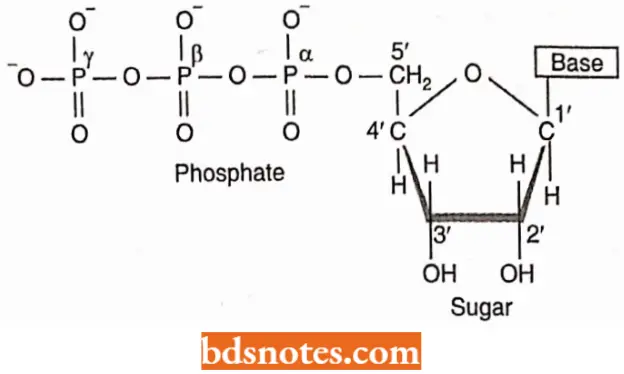
- The precursors in the synthesis of RNA are the four ribonucleotide 5’- triphosphates (rNTP)-ATP, GTP, CTP, and UTP.
- On the ribose portion of each NTP, there are two OH groups – one each on the 2’-and 3’-carbon atoms.
- In the polymerization reaction, a 3’- OH group of one nucleotide reacts with the 5’-triphosphate of a second nucleotide; a pyrophosphate is removed, and a phosphodiester bond results.
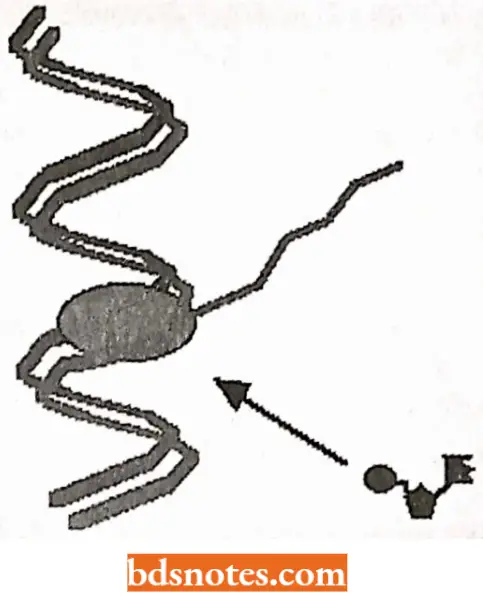
- The sequence of bases in RNA molecules is determined by the base sequence of the DNA. Each base, added to the growing end of the RNA chain, is chosen by its ability to base-pair with the DNA strand used as a template; thus, the bases C, T, G, and A in a DNA strand cause G, A, C, and U respectively, to appear in the newly synthesized RNA molecule.
“Importance of studying transcription stages for biology students: Questions explained”
- The DNA molecule being transcribed is double-stranded, yet in many particular regions, only one strand serves as a template.
- The RNA chain grows in the 5’→ 3′ direction; that is, nucleotides are added only to the 3’- OH end of the growing chain. (This is the same as the direction of chain growth in DNA synthesis).
- The RNA molecule is terminated by a 5′-triphosphate at the non-growing end.
- The RNA molecule is antiparallel to the DNA strand being copied.
- Once initiated, RNA chains grow at a rapid rate – 40 to 50 nucleotides per second in E.coli at 37°C.
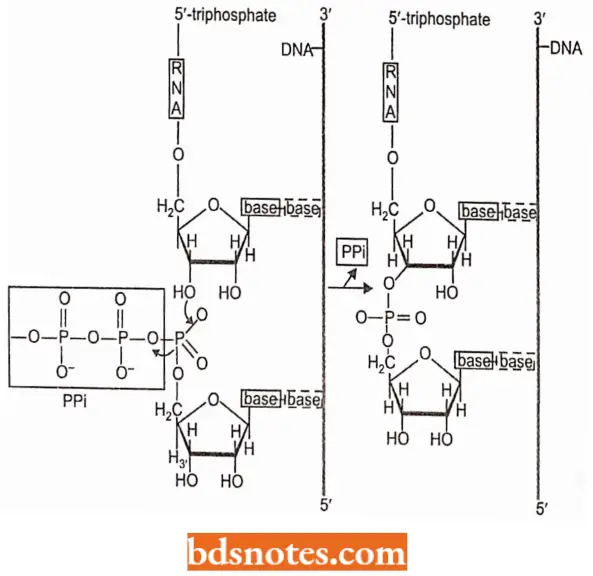
- RNA polymerases, in contrast with ON A polymerases, me able to initiate chain growth; that is, no primer is needed.
- Only ribonucleoside 5′-triphosphates participate in RNA synthesis and the first base to be laid down in the initiation event is a triphosphate.
- Its 3′-OH group is the point of attachment of the subsequent nucleotide. Thus, the 5′ end of a growing RNA molecule terminates with a triphosphate.
The Overall Polymerization Reaction Can Be Written As Follows:
![]()
- Which XTP represents the first nucleotide at the 5′ terminus of the RNA chain, NMP is a mononucleotide in the RNA polymerase, and PP1 is the pyrophosphate released each time a nucleotide is added to the growing chain.
- The Mg2+ is required for all nucleic acid polymerization reactions.
The RNA Polymerase Enzyme: In E. coli and other prokaryotes, a single RNA polymerase (RNA-P) enzyme is responsible for the synthesis of all kinds of RNAs (such as mRNA, tRNA, and rRNA).
- RNA-P is, in fact, one of the largest enzymes known (MW 490,000).
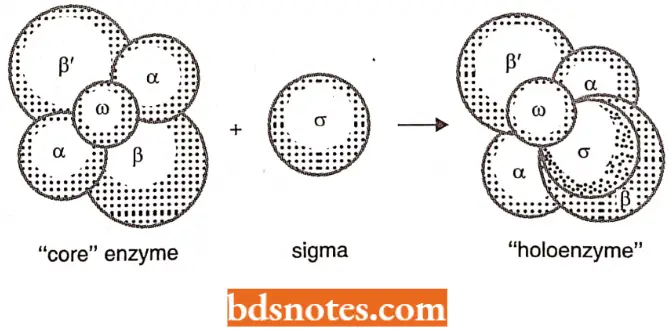
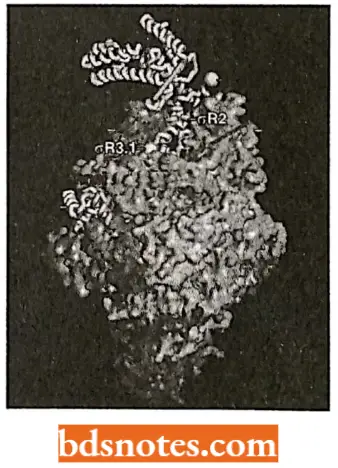
- It consists of six subunits (i.e., polypeptide chains)-two identical alpha (α) subunits and one chain of each of beta (β), bete dash (β’), omega (ω), and sigma (σ) subunits.
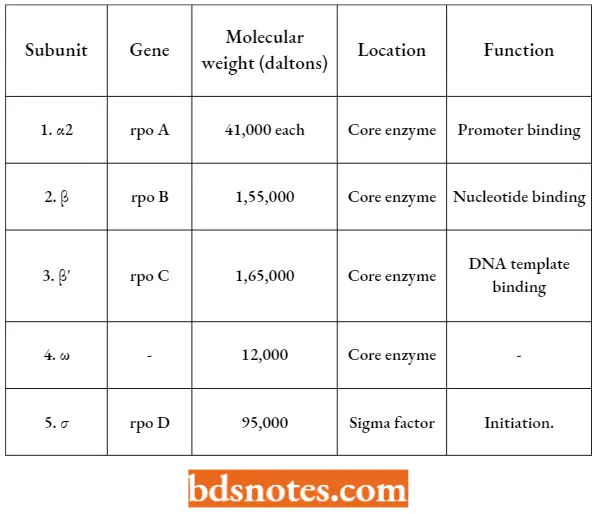
- Some characteristics of each subunit of RNA-P have been tabulated.

- The complete RNA polymerase enzyme is termed holoenzyme and can be represented as α2ββ’ωσ in which attachment of sigma (α) subunit (or factor) is not very firm, but the resultant core enzyme (α2ββ’) does not lose its catalytic activity of transcription. The active sites of the core enzyme.
- Functions of different polypeptide subunits of RNA polymerase are now known, but not in any detail.
- Thus, β and β’ subunits form the catalytic center of RNA-P and help RNA polymerase in the unwinding of DNA molecules for transcription.
- The sigma (σ) factor helps in the recognition of start signals on DNA molecules and directs RNA polymerase in selecting the initiation sites (promotor).
- Once RNA synthesis is initiated and the RNA molecule becomes 8-9 bases long, the factor dissociates from the holoenzyme, and then the core enzyme brings about the elongation of mRNA (or any other RNA).
“Common challenges in understanding transcription stages effectively: FAQs provided”
- Binding of RNA Polymerase to Promoter and Initiation Unlike replication, transcription does not progress along the entire length of a chromosome. Instead, certain parts of the chromosome arc are transcribed.
- Only one of the two strands of a DNA duplex is transcribed; this strand is called the sense strand.
- The other strand of DNA which is not transcribed at the moment, is called the anti-sense strand. Like DNA.
- RNA is synthesized in the 5 → 3′ direction. Active centers in the core enzyme from the single-stranded region of the bacterial RNA polymerase enzyme.
- DNA template. This localized unwinding moves along the molecule followed by single-recoiling of the helix behind the newly synthesized RNA.
- The region of the sense strand of DNA which is transcribed into RNA, is called the coding region.
Some Basic Characteristics Of Different Subunits (=Polypeptides) Of RNA Polymerase Of E.coli:
- The DNA region that the RNA polymerase enzyme associates with immediately before beginning transcription is known as the promoter.
- The promoter is a significant part of gene expression in both prokaryotes and eukaryotes.
- Promoters contain the information for transcription initiation and are the major sites in which gene expression is regulated.
- The molecule of enzyme of RNA polymerase covers a region of about sixty base pairs of DNA.
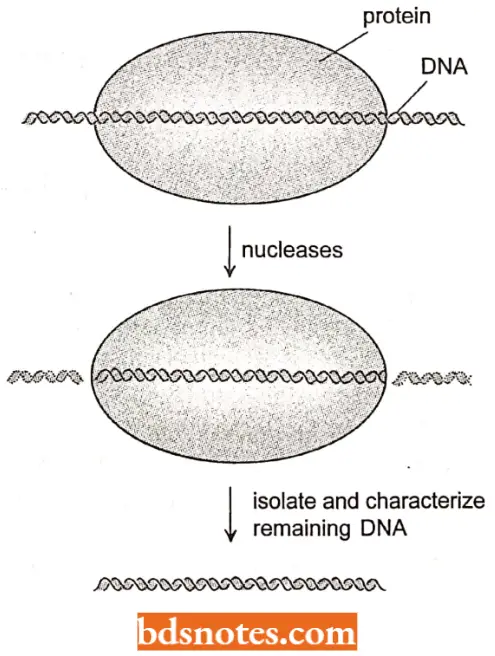
- This point was determined by causing the polymerase to bind to DNA and then digesting the mixture with nuclease enzymes, in a technique known as footprinting.
- The polymerase “protects” or prevents degradation of the region it covers. The undigested DNA is then isolated and its size is determined.
- Geneticists have gained much new information about the nature of recognition regions within promoters through recombinant DNA technology and nucleotide sequencing techniques.
- Sequencing of numerous promoters has revealed that they contain common sequences.
- If the promoter nucleotide sequences align with each other, and each has the same series of nucleotides in a given segment, then a sequence of that segment is regarded as a conserved sequence.
- If, however, there is some variation in the sequence, but certain nucleotides occur at a high frequency (significantly greater than by chance), then these nucleotides are said to form a consensus sequence.
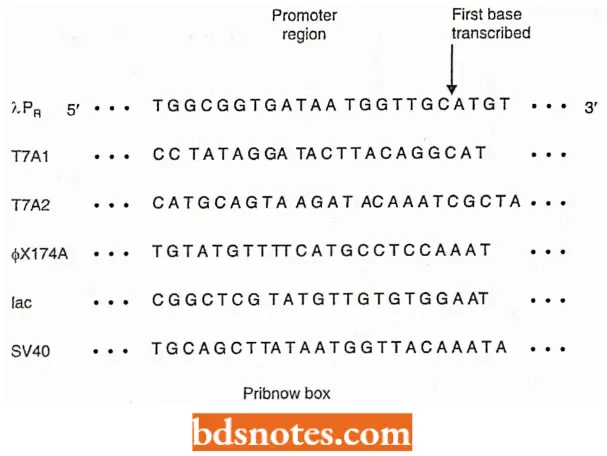
Surrounding a point in prokaryotic promoters about ten nucleotides before the first transcribed base is just such a consensus sequence – TATAAT. This sequence is called the Pribnow box after one of its discoverers.
- The nucleotides in the Pribnow box are mostly adenines and thymines (T), so the region is primarily held together by only two hydrogen bonds per base pair.
- Since local DNA denaturation occurs during transcription by RNA polymerase (i.e., the DNA is opened to allow transcription), fewer hydrogen bonds make this process easier energetically.
- When the RNA polymerase is bound at the promoter region, it is in position to begin polymerization of six to eight nucleotides down from the Pribnow box.
- I have been shown the sequences which represent the coding strand of DNA.
- It is general convention to show the coding strand because both that strand and mRNA have the same sequences, substituting U for T in RNA, they are both complementary to the same template strand (which is also known as anticoding strand or noncoding strand’,.
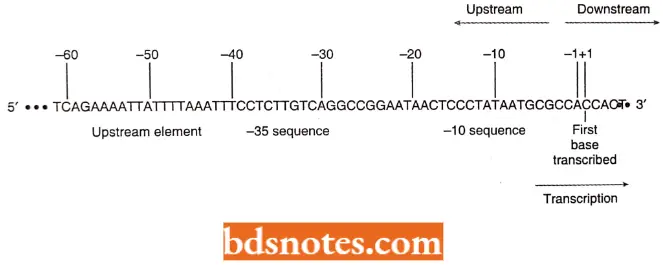
- Another convention is to indicate the first base transcribed by the number +1 and to use positive numbers to count farther down the DNA in the downstream direction of transcription.
“Why is early learning of transcription critical for molecular biology? Answered”
- If transcription is proceeding to the right, the direction to the left is called upstream, with bases indicated by negative numbers Under this convention, the Pribnow box is often referred to as the -10 sequence.
- Indicates another region with similar sequences among many promoters centered near -35 and referred to as the – 35 sequence. The consensus sequence at -35 is TTGTCA.
- Mutation studies have tried to determine the relative roles of the -10 and -35 sequences in transcription.
- In other words, mutations of bases in the -10 and -35 regions of the promoter were examined to determine how they affected transcription initiation.
- The conclusion from these studies is that both regions contribute to the efficiency of binding of polymerase enzyme to the promoter. In other words, the more each sequence differs from the consensus sequence, the less frequently that promoter initiates the transcription.
- The sigma factor recognizes both the -35 and the -10 sequences.
- The sigma factor is also sensitive to the spacing between these sequences, preferring (i.e., being most efficient at seventeen base pairs.

- Moreover, in bacterial chromosomes, farther upstream from the – 35 sequence is a recognition element that is very strongly expressed, for example, ribosomal RNA genes. this upstream element or UP element; is about twenty base pairs long, is centered at -50, and is rich in A and T.
- By mutational studies, it has been shown that adding the UP element to promoters that do not normally have it greatly increases the rate of transcription.
- There are other recognition sites in prokaryotes, both upstream and downstream, at which various proteins attach that can enhance or inhibit transcription by direct contact with the polymerase (the a and subunits).
- Because the holoenzyme (RNA polymerase) recognizes consensus sequences in a promoter, it is not surprising that some promoters are bound more efficiently than others or that different sigma factors exist within a cell.
- For example, in E. coli, the major sigma factor is a protein of 70,000 daltons, referred to as O70.
- There exist about five less common sigma factors, due to which a bacterial cell remains able to transcribe different genes under different circumstances.
- For instance, in an E.coli cell subjected to elevated temperatures, a group of new proteins, called heat shock proteins, protect the cell to some extent against the elevated temperatures.
- These heat shock proteins all appear at once because they have promoters that a different sigma factor recognizes, one with a molecular weight of 32,000 daltons (σ32); this new sigma factor is produced by the cell after heat shock.
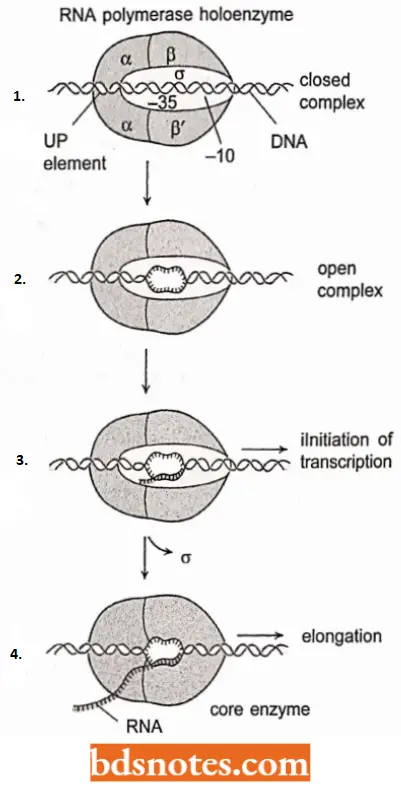
- According to the current picture of the mode of action of polymerase holoenzyme, this enzyme sets down on a DNA promoter because the sigma factor recognizes the -10 and – 35 elements, the α proteins recognize the UP element and the α and σ subunits recognize proteins to bound to various other upstream elements, when present.
- This initiation complex is initially known as a closed complex because the DNA has not melted, which is the next step in transcription initiation. After the transcription of 5 to 10 bases, the sigma factor is released.
- Once an open promoter complex has formed, RNA polymerase is ready to initiate RNA synthesis. RNA polymerase contains two nucleotide-binding sites, called the initiation site and the elongation site.
- The initiation site binds only purine triphosphates, namely ATP and GTP, and one of these (usually ATP) is the first nucleotide in the growing RNA chain.
“Factors influencing success with transcription knowledge: Q&A”
- Thus, the first DNA base that is transcribed is usually thymine(T).
- The initiating nucleoside triphosphate binds to the enzyme in the open-promoter complex and forms a hydrogen bond with the complementary DNA base.
- The elongation site is then filled with a nucleoside triphosphate that is selected strictly by its ability to form a hydrogen bond with the next base in the DNA strand.
- The two nucleotides are then joined together, the first base is released from the initiation site, and initiation is completed.
- The dinucleotide remains hydrogen-bonded to the DNA.
- The elongation phase begins when the polymerase releases the base and then moves along the DNA chain.
Elongation phase: Since there is just one bacterial RNA polymerase, the general mechanism of transcription is the same for all bacterial genes.
- The following description of elongation and termination, given in the context of mRNA synthesis, therefore applies equally well to the synthesis of non-coding RNA (i.e., rRNA and tRNA).
- During transcription, ribonucleotides are added one after another to the growing 3′ end of the RNA transcript, the identity of each nucleotide specified by the base-pairing rules: A base pairs with T or U; G base pairs with C.
- During each nucleotide addition, the β- and γ- phosphates are removed from the incoming nucleotide, and the hydroxyl group is removed from the 3′ – carbon of the nucleotide present at the end of the chain.
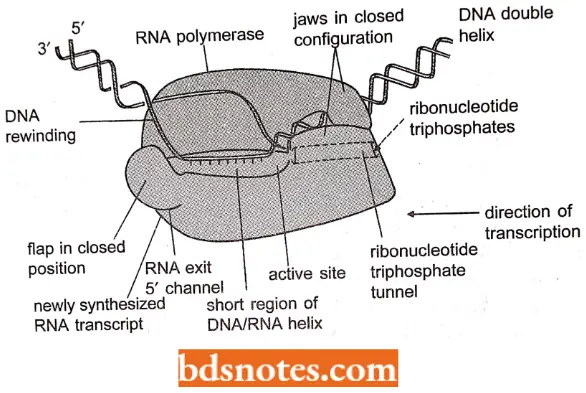
- At this stage of transcription, the bacterial RNA polymerase is in its core enzyme form, comprising four proteins, two relatively small (approximately 35 k Da) α subunits, and one each of the related subunits β and β’ (both approximately 150 k Da): the subunit that has the key role in initiation has now left the complex.
- The RNA polymerase covers about 30 bp of the template DNA, including the transcription bubble of 12-14 bp, within which the growing transcript is held to the template strand of the DNA by approximately eight RNA-DNA base pairs.
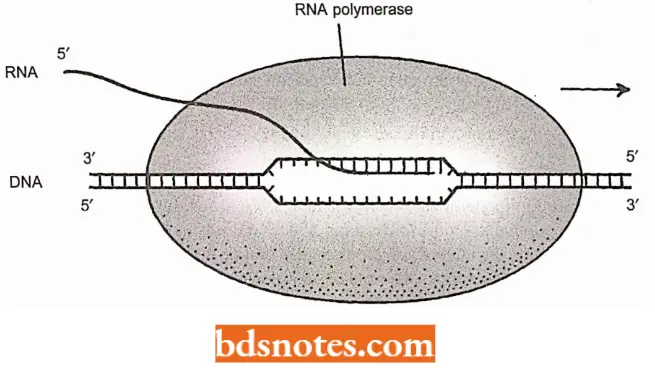
- The polymerase has to keep a tight grip on both the DNA template and the RNA that it is making to prevent the transcription complex from falling apart before the end of the gene is reached.
- However, this grip must not be so tight as to prevent the polymerase from moving along the DNA.
- In the current model, which has been produced by X-ray crystallographic studies and crosslinking experiments, the DNA template lies between the β and β’ subunits, within a trough on the enclosed surface of β’.
- The active site for RNA synthesis also lies between these two subunits, with the non-template strand of DNA held within the β subunit and RNA transcript extruded from the complex via a channel formed partly by the β and partly by the β’ subunit (Korzheva et ah 2000).
Termination Phase: The exact mechanism involved in the termination of transcription is still not known.
- However, molecular biologists believe that transcription is a stepwise addition process of nucleotide-by-nucleotide, with the polymerase pausing at each position and making a choice between continuing elongation by adding another ribonucleotide to the file transcript or terminating by dissociating from the template.
- Which choice is selected depends on which alternative is more favorable in thermodynamic terms (Von Hippcl, 1998).
- This model emphasizes that for termination to occur, the polymerase has to reach a position on the template where dissociation is more favorable than continued RNA synthesis.
“Steps to explain the stages of transcription: Initiation vs elongation vs termination: Q&A guide”
The current view holds that bacteria appear to use the following two distinct schemes for transcription termination.
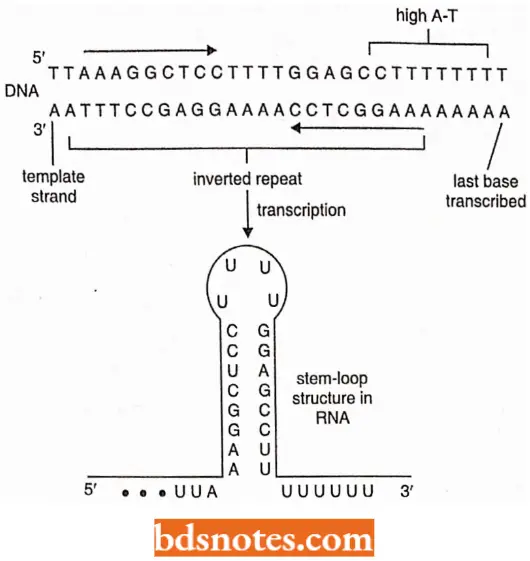
Transcription termination at an intrinsic terminator: About half the positions in E. coli at which transcription terminates correspond to DNA sequences where the template strand contains an inverted palindrome followed by a run of deoxyadenosine nucleotide.
- These intrinsic terminators have been thought to promote dissociation of polymerase by destabilizing the attachment of the growing transcript to the template, in two ways.
- First, when the inverted palindrome is transcribed, the RNA sequence folds into a stable hairpin (the nascent RNA hairpin; this RNA – RNA base pairing is favored over the DNA – RNA pairing that normally occurs in the transcription bubble.
- This reduces the number of contacts made between the template and transcript, weakening the overall interaction and favoring dissociation.
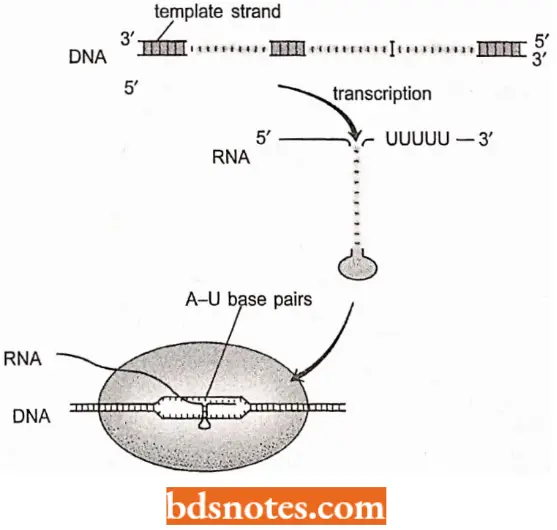
- The interaction is further weakened when the run of As (= adenine bases) in the template is transcribed, because the resulting A-U base pairs have only two hydrogen bonds each, compared with three for each G-C pair.
- The net result is that termination is favored over continued elongation (Von Hippel, 1998).
- An alternative model which is based on crosslinking experiments has shown that the RNA hairpins make contact with flap structure on the outer surface of the RNA polymerase (β subunit, adjacent to the exit point of the channel through which the RNA emerges from the complex.
- Although the flap structure is quite distant (some 6.4 nm) from the active site of the polymerase, a direct connection is made between the two by a segment of β-sheet within the β-subunit.
- Movement of the flap could therefore affect the positioning of amino acids within the active site, possibly leading to breakage of the DNA-RNA base pairs and termination of transcription.
- Toulokhonov et al., have demonstrated the presence of a protein called NusA which enhances termination at intrinsic promoters, interacts with the hairpin loops and flap structure, and may stabilize the contact between the two.
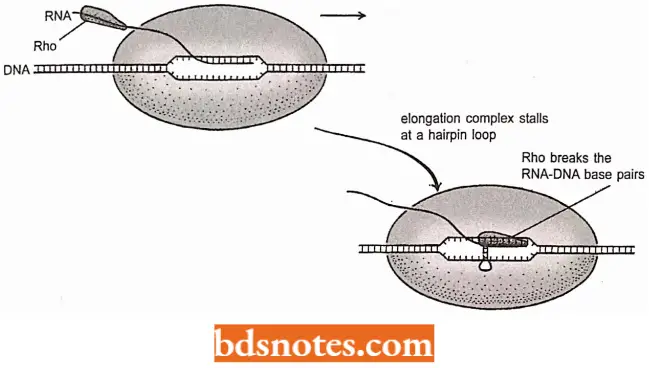
- Rho-dependent transcription termination. The second type of bacterial termination signal is Rho-dependent.
- These signals usually retain the hairpin feature of intrinsic terminators, although the hairpin is less stable and there is no run of As in the template.
- Termination requires the activity of a protein called Rho, which attaches to the transcript and moves along the RNA toward the polymerase.
- If the polymerase continues to synthesize RNA then it keeps ahead of the pursuing Rho, but at the termination signal the polymerase stalls.
- Rho is a helicase, which means that it actively breaks base pairs, in this case between the template and transcript, resulting in the termination of transcription.
“Role of RNA polymerase in synthesizing RNA from DNA: Questions answered”
Antitermination and attenuation: In bacteria, the following two mechanisms have evolved to influence the repeated choice that the polymerase has to make between elongation and termination when copying a template (DNA).
Antitermination: This process occurs when the RNA polymerase ignores a termination signal and continues elongating its transcript until a second signal is reached.
Antitermination provides a mechanism whereby one or more of the genes at the end of an operon can be switched off or on by the polymerase recognizing or not recognizing a termination signal located upstream of those genes.
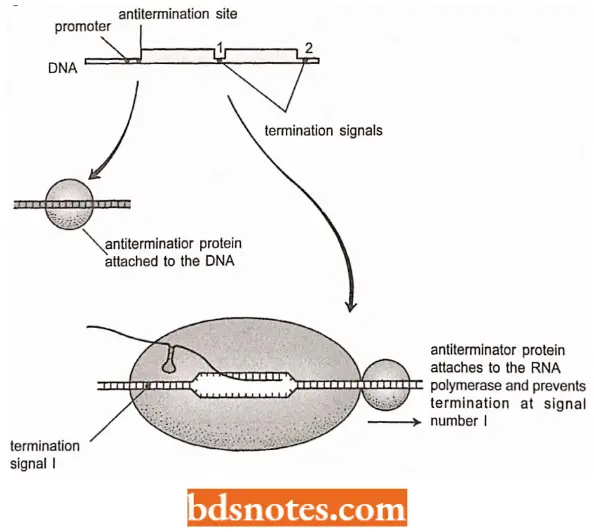
- Antitermination is controlled by an antitermination protein (for example N protein and Q protein of the genome of bacteriophage X of E.coli; Friedman et al., 1987).
- This protein attaches to the DNA near the beginning of the operon and then transfers to the RNA polymerase as it moves past en route to the first termination signal.
- The presence of the antitermination protein causes the enzyme to ignore the termination signal, presumably by countering the destabilizing properties of an intrinsic terminator or by preventing stalling at a Rho-dependent terminator.
- Although the mechanics of the process are unclear, the impact that antitermination can have on gene expression has been described in detail, especially during the infection cycle of bacteriophage X (Friedman et al., 1987).
Attenuation: The process of attenuation operates primarily with operons that code for enzymes involved in amino acid biosynthesis. The tryptophan operon of E. coli shows how it works.
- In the tryptophan operon, two hairpin loops can form in the region between the start of the transcript and the beginning of trpE.
- The smaller of these loops acts as a termination signal, but the larger hairpin loop, which is closer to the start of the transcript, is more stable.
- The larger loop overlaps with the smaller loop (i.e., termination hairpin), so only one of the two hairpins can form at any one time.
- Which loop forms depends on the relative positioning between the RNA polymerase and a ribosome which attaches to the 5′ end of the transcript as soon as it is synthesized to translate the genes into protein.
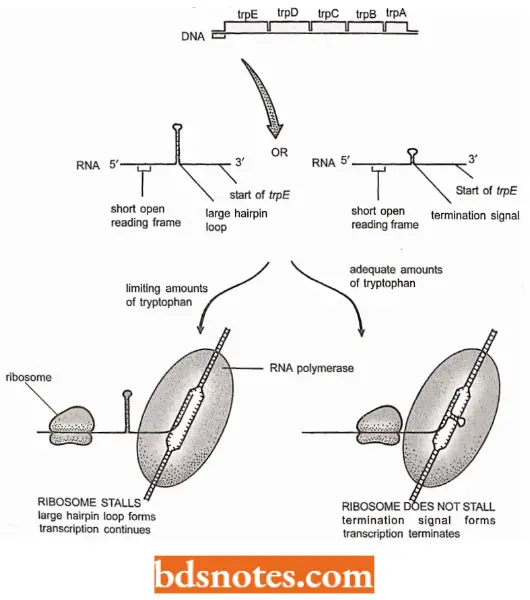
- Thus, if the ribosome stalls so that it does not keep up with the polymerase, then the larger hairpin forms and transcription continues.
- However, if the ribosome keeps pace with the RNA polymerase then it dislocates the larger hairpin by attaching to the RNA that forms part of the stem of this hairpin When this happens the termination hairpin can form, and transcription stops.
“How do promoters and terminators regulate transcription? FAQ explained”
- Stalling of the ribosome can occur because upstream of the termination signal is a short open reading frame (ORF) coding for a 14-amino-acid peptide that includes two tryptophans.
- Again, if the amount of free tryptophan is limited, then the ribosome stalls as it attempts to synthesize this peptide, while the polymerase continues, to make its transcript (mRNA).
- Because this transcript contains copies of genes coding for the biosynthesis of tryptophan, its continued elongation undertakes the requirement that the cell has for this amino acid.
- When the amount of tryptophan in the cell reaches a satisfactory level, the attenuation system prevents further transcription of the tryptophan operon, because now the ribosome does not stall while making the short peptide, and instead keep pace with the polymerase, allowing the termination signal to form.
- Tryptophan operon of E.coli is controlled not only by attenuation but also by a repressor.
- Exactly how attenuation and repression work together to regulate expression of the operon is not known, but it is believed that repression provides the basic on-off switch and attenuation modulates the exact level of gene expression that occurs.
- Other operons of E.coli, such as those for the biosynthesis of histidine, leucine, and threonine, are controlled entirely by attenuation.
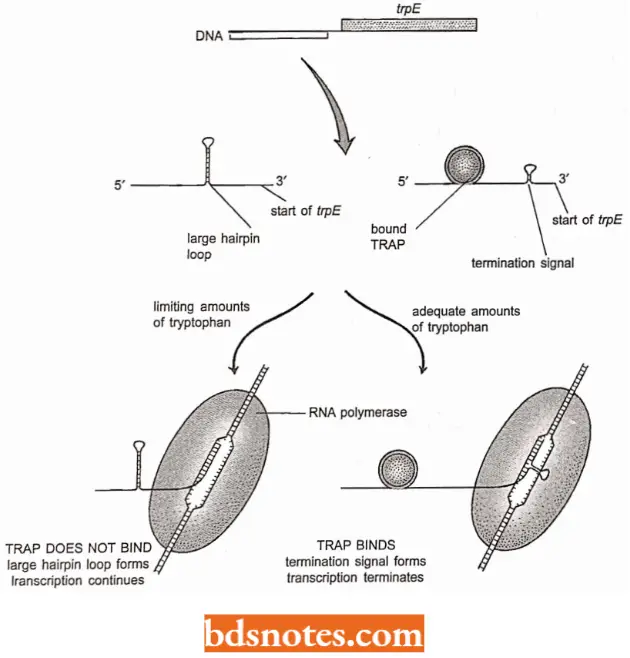
- Interestingly, in some bacteria, including Bacillus subtilis, the tryptophan operon is one of those that is regulated entirely by attenuation.
- In these bacteria, attenuation is mediated not by the speed at which the ribosome tracks along the mRNA, but by an RNA-binding protein called trp RNA-binding attenuation protein (TRAP) which in the presence of tryptophan, attaches to the mRNA in the region equivalent to the short ORE of E.coli transcript.
- The attachment of TRAP leads to the formation of the termination signal and stoppage of transcription (Antson et al., 1999).
Initiation Of Transcription In Eukaryotes
The mRNA molecules make up the transcriptosome (i.e., the entire mRNA content of a cell) and specify the protein content of the cell.
- As the central players in genome expression, mRNAs have received the greatest attention from molecular biologists.
- Transcriptional events in prokaryotes (bacteria) are different in many respects from those in eukaryotes. In bacteria, mRNA does not undergo any significant forms of processing: the primary transcript that is synthesized by the RNA polymerase in itself is the nature of mRNA, and its translation usually begins before transcription is complete.
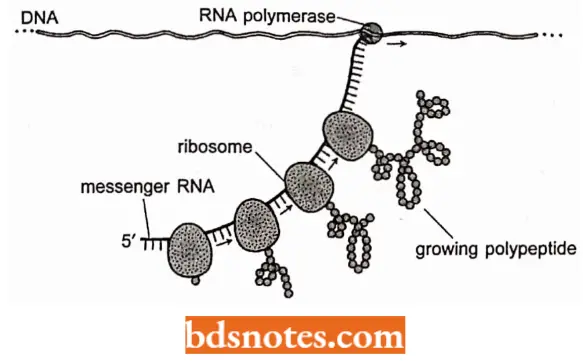
- The messenger RNA is synthesized in the 5′ → 3′ direction and it is near the 5′ end of mRNA that translation begins. As soon as the 5′ end of mRNA is available, a ribosome can attach to the mRNA and move along it in the 5′ → 3′ direction lengthening the growing polypeptide as it moves.
- When the first ribosome moves away from the 5″ end of the transcript, a second ribosome can attach and begin translation.
- These processes are repetitive. In eukaryotes, however, messenger RNA is synthesized in the nucleus, but protein synthesis takes place in the cytoplasm.
- (Such a regional division of labor is not present in E. coli because the bacterium has no nucleus). Before an mRNA leaves the nucleus, it is highly modified by processes that generally do not occur in prokaryotes.
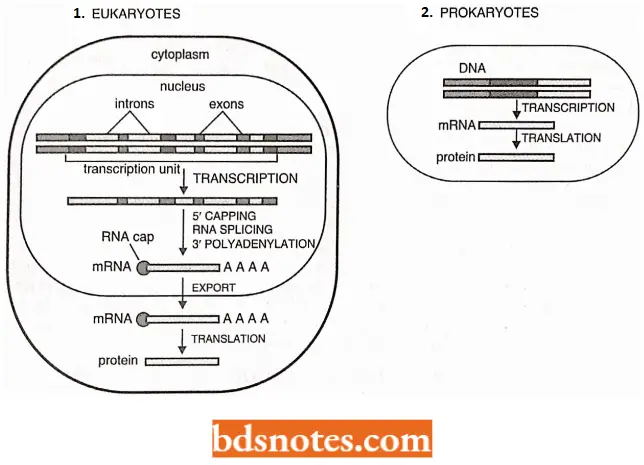
Eukaryotic processing of primary transcript includes:
- 5’ capping;
- RNA splicing and
- 3’ polyadenylation.
Here now we will describe the initiation of transcription in eukaryotes.
“Early warning signs of gaps in understanding transcription basics: Common questions”
Promoters: Eukaryotic promoters are somewhat similar to prokaryotic promoters; both are regions of ON A at the beginning of genes with signals that allow RNA polymerase to attach and begin transcription.
In eukaryotes, however, more proteins are involved in promoter recognition, and more proteins are involved in the control of transcription, many recognizing signals thousands of base pairs away.
- In eukaryotes, there are three major classes of RNA polymerases which are designated as 1, 2, and 3, and are found in the nucleus.
- The three polymerases have different properties and can be distinguished by the ions required for their activity, the optimal ion strength, and their sensitivity to inhibition by various antibiotics (for example., α- amanitin).
- Each of these enzymes is a large protein (˜500,00 daltons), with two large and several (8 to 10) smaller subunits.
Properties Of Three Different Eukaryotic Nuclear Rna Polymerases:
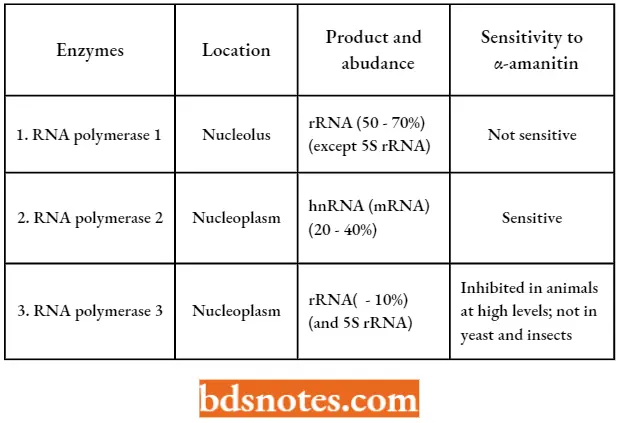
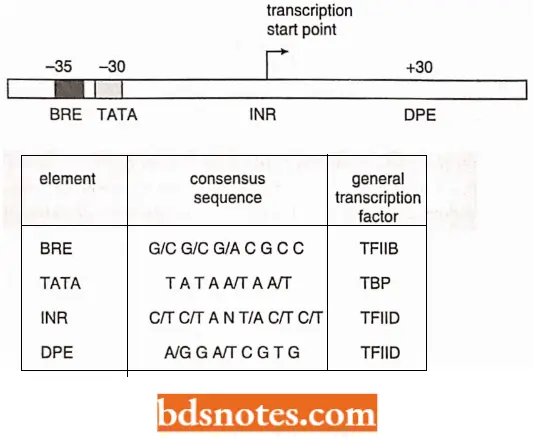
- All three eukaryotic RNA polymerases (1, 2, and 3) recognize a seven-base sequence, TATAAAA. located at about -25 on the promoter DNA.
- It is similar to the -10 sequence in prokaryotes and is called the TATA box (or Hogness box after it’s discovered, D. Hogness). Since RNA polymerase II transcribes most of the genes in eukaryotes, so let us describe it in sufficient detail.
- Among the large number of promoters that have been sequenced, a few lack the TATA box, yet are still transcribed.
- Transcription initiation in these promoters appears to be controlled by a CT-rich area, called the initiator element (INR), at +1 of the transcript (close to the transcription start site), coupled with a downstream promoter element (DPE) at about +28 to the +34 of the transcript.
“Asymptomatic vs symptomatic effects of ignoring transcription principles: Q&A”
- In TATA-less promoters, a protein called TFIID requires both these elements to bind.
- The initiator element has a consensus sequence of TCAGTT or TCATTC, and the downstream promoter element has a consensus sequence of AGACGTG or GGTCGTG.
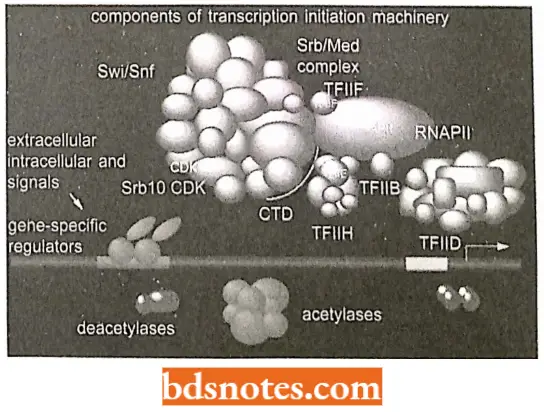
General Transcription Factors: RNA polymerase II of yeast a protein of twelve subunits, enzyme cannot locate RNA Polymerase Holoenzyme.
- Promoters or stably attach to DNA. To attach at the beginnings of genes, RNA polymerase II must interact with several proteins called general transcription factors.
- The proteins are “general” because they assemble on all promoters used by RNA polymerase II. In eukaryotes, general transcription factors are named after the polymerase they work with.
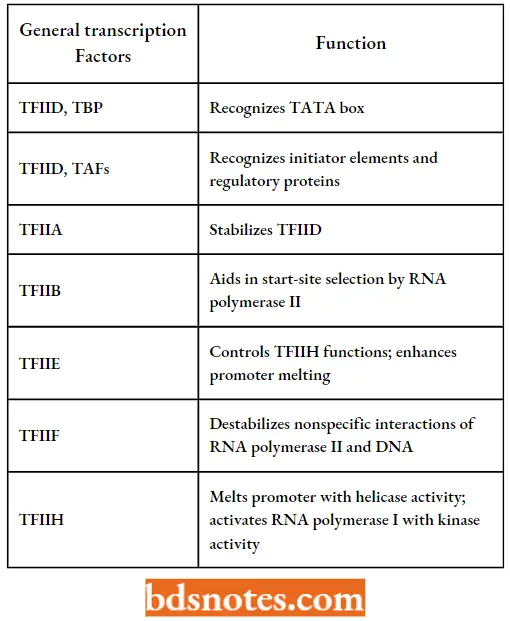
- Thus, the transcription factor that recognizes the TATA box for polymerase II genes is called TFIID (D being the fourth letter of the alphabet for the fourth transcription factor so named).
TFIID is composed of one subunit that recognizes the TATA sequence, called TATA-binding protein (TBP), and upto a dozen other proteins called TBP-associated factors (TAFs), which recognize the initiator element, when present and aid in regulating transcription.
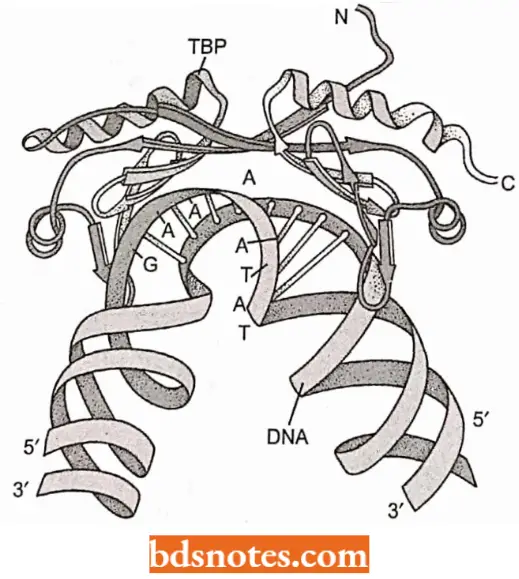
- TFIID is virtually similar to the sigma factors of prokaryotic RNA polymerase. One interesting aspect of the binding of TBP is that it causes a significant bending and opening of the DNA.
- This bending may be an important signal for other binding proteins.
Techniques Employed In the Study Of Initiation Of Eukaryotic Transcription: Much of the information regarding the initiation of transcription in eukaryotes has been gathered by footprinting, mutational studies, cloning and isolating the genes and proteins involved, and then reconstituting various purified combinations in test tubes.
These studies have been combined with kinetic research to determine which arrangements are stable, immunological research to isolate various components with antibodies, and photocrosslinking studies to determine which moieties are in contact with each other.
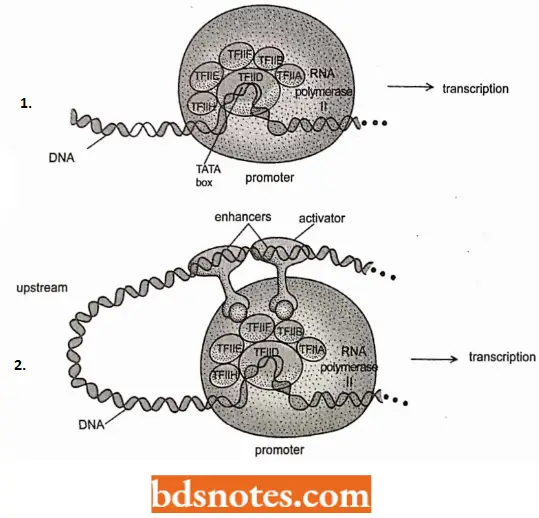
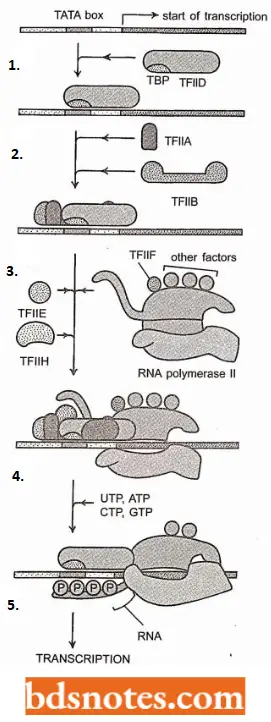
- Once TFIID binds to the TATA box, a cascade of recruitment (i.e., binding) of other transcription factors takes place. Transcription factors TFIIA, TFIIB, and TFIIF bind, as does RNA polymerase II in an unphosphorylated state.
- In the next stage, transcription factors TFIIE and TFIIH bind, forming a pre-initiation complex (PIC), equivalent to the E. coli holoenzyme.
- The RNA polymerase II is then phosphorylated, presumably by TFIIH, which is a kinase; at this point, most of the transcription factors drop off, leaving the elongation complex, which carries out a basal rate of transcription.
“Can targeted interventions improve outcomes using transcription knowledge? FAQs provided”
TFIIH also has a role here since it is a helicase that summarizes the postulated roles of the general transcription factors.
Postulated Roles Of The General Transcription Factors Of Rna Polymerase 2 (Source Tamarin, 2002):
Phosphorylation Of C-Terminal Domain: Like the bacterial polymerase, polymerase 2 remains at the promoter synthesizing short lengths of RNA until it undergoes a conformational change and is released to begin transcribing a gene.
- A key step in this release is the addition of phosphate groups to the “tail” of the RNA polymerase (which is known as the CTD or C-terminal domain).
- This phosphorylation is also catalyzed by TFIIH, which in addition to a helicase contains a protein kinase as one of its subunits.
- The polymerase can then disengage from the cluster of general transcription factors, undergoing a series of conformational changes that tighten its interaction with DNA and acquire new proteins that allow it to transcribe for long distances without dissociating.
- Specific Transcription Factors. For activated transcription, a high level of transcription, to take place, other factors are needed that are involved in controlling which promoters are actively transcribed.
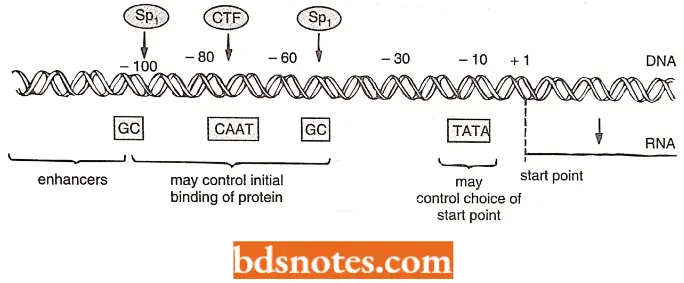
- These other factors are activators or specific transcription factors that can bind to DNA sequences, called enhancers. Enhanccra is often hundreds or thousands of base pairs upstream from the promoter.
- These specific transcriptional activators have domains (regions) that recognize their specific enhancer sequences, regions dial recognize proteins associated with the polymerase (general transcription factors), and regions that allow the joint attachment of other transcription factors.
- Similar to activators and enhancers, repressors can bind to silencer regions of DNA, often far upstream of the promoters, to repress transcription.
- Thus, many genes are associated with numerous and complex arrangements of transcription factors, providing intricate control of transcription.
- For specific transcription factors to attach to both enhancers and the polymerase machinery, possibly thousands of base pairs apart, the DNA bends to allow them to come into the range of polymers.
“Difference between prokaryotic and eukaryotic transcription: Q&A explained”
- Although polymerase 1 and 2 seem to have termination signals similar to rho-independent promoters in prokaryotes, termination of transcription of RNA polymerase 2 genes is more complex, coupled with further processing of the mRNA.
- Further unlike prokaryotic RNA polymerases, eukaryotic RNA polymerases do proofreading (showing 3′ → 5′ exonuclease activity).
- Moreover, eukaryotic DNA is complex with histone proteins that can interfere with transcription. In turn, part of the RNA polymerase 2 complex is made up of proteins that can disrupt the histones bound to the DNA.
In addition, the RNA polymerase 2 complex contains proteins that act as mediators between activators and the polymerase holoenzyme.
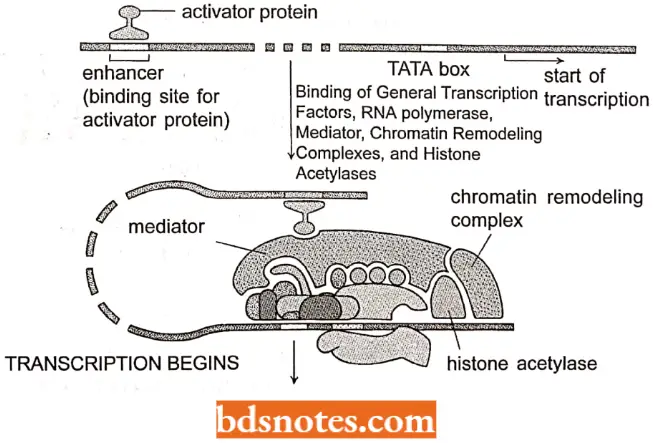
- This complex coordination of the initiation of transcription in eukaryotes has been termed combinatorial control, the huge initiation complex may contain 100 or more different polypeptides.
- Finally, transcription in archaea (archaebacteria), although under much simpler control than in eukaryotes, resembles transcription in eukaryotes rather than prokaryotes.
- The study of details of the transcription process initiation, control, and termination – is one of the most active and exciting areas in modem genetics.
Control Of Transcription In Eukaryotes
In prokaryotes, an RNA polymerase holoenzyme with its promoter-recognizing sigma factor is generally active, transcribing at high levels; repressors are needed to prevent transcription.
In eukaryotes, an RNA polymerase holoenzyme (for example., RNA polymerase 2), with its recognizing TFIID, is generally not transcribing; it needs access to the promoter, which is usually wrapped around nucleosomes, and it needs specific transcription factors to become active.
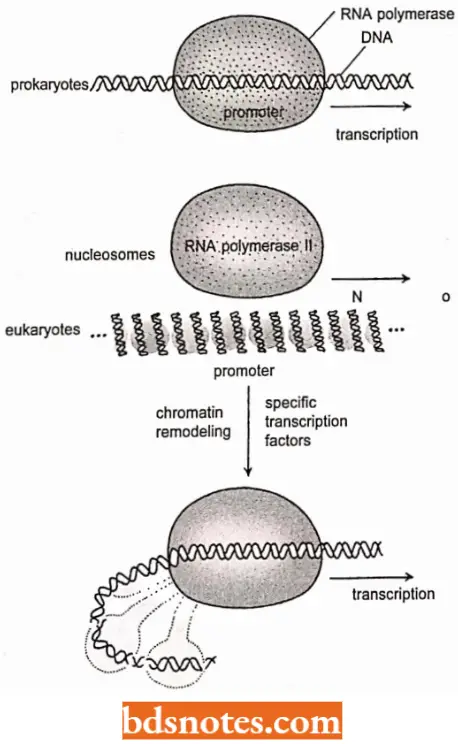
- Thus, although the parts of the transcribing machinery of prokaryotes and eukaryotes are generally similar, the essence of prokaryotic transcription is activity, whereas the essence of eukaryotic transcription is inactivity.
- In addition, eukaryotes generally do not have onerous; however, groups of eukaryotic genes involved in the same pathway or function can be induced simultaneously by having common enhancers that respond to the same specific transcription factors. Such a group of genes is called a synexpression group.
Chromatin remodeling: For transcription to take place in eukaryotes the DXA must be available for the preinitiation complex to form, with its RNA polymerase and genera) transcription factors.
- It appears that DNA wrapped around nucleosomes is often not accessible for the formation of the preinitiation complex, but is available for recognition by transcription-activating proteins, also called specific transcription factors (as compared to general transcription factors of the RNA polymerase machinery).
- One model of initiation of transcription by genes whose promoters are wrapped around nucleosomes is for specific transcription factors to recruit chromatin-remodeling proteins.
- There are two general classes of proteins that remodel nucleosomes: histone acetyltransferases and ATP-dependent chromatin remodeling proteins such as the SWI/SNF complex in yeast.
- Thus, the presence of one or more specific transcription factors can begin the process of transcription by recruiting chromatin-remodeling proteins that allow the RNA polymerase access to the promoter.
Specific transcription factors: Eukaryotic transcription starts with the formation of a preinitiation complex formed by the amalgamation of a group of general transcription factors (such as TFIJD in RNA polymerase II formation).
- Proteins that exert control over transcription at specific promoters are the specific transcription factors. These proteins generally have two domains: a domain that recognizes specific DNA sequences, and a domain that recognizes another protein such as a protein in the preinitiation complex.
- Thus, these proteins recognize signals in the vicinity of the promoter of a gene, bind there, and initiate transcription. Currently, we believe that the majority of specific transcription factors act by recruiting the components of the DNA polymerase holoenzyme.
- Thus, the binding of a specific transcription factor at a promoter is the first step in the formation of a pre-initiation complex at the promoter of a gene.

“Most common complications of poorly understood transcription concepts: FAQs”
- Some transcription-activating proteins also recruit chromatin-remodeling proteins.
- An example of a specific transcription factor is the Dorsal protein, the product of the dorsal gene in fruit flies, active in development.
- Dorsal protein controls the transcription of several genes and at several different levels of protein concentration.
- The ability to have different effects at different concentrations is extremely important, allowing gradients of the same protein to control the expression of different genes.
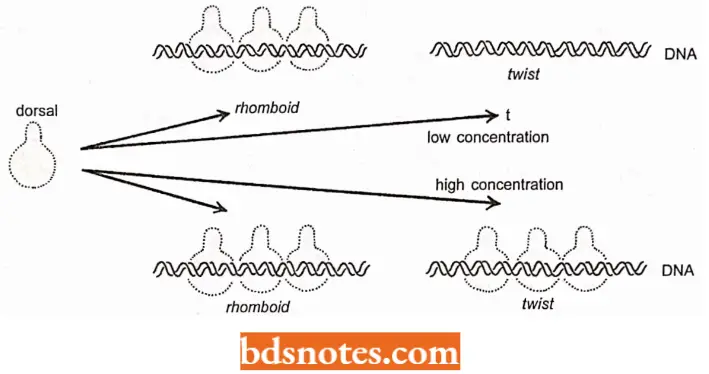
- One gene Dorsal protein controls is the rhomboid, which has three sites in its promoter that the Dorsal protein binds to, initiating transcription.
- Another gene, twist, also has three sites in its promoter that bind the Dorsal protein, initiating transcription.
- However, the rhomboid sites are more efficient in binding Dorsal protein; thus, rhomboid is transcribed at lower concentrations of Dorsal protein than twist is.
- One other signal in the control of transcription that is of current interest is methylation.
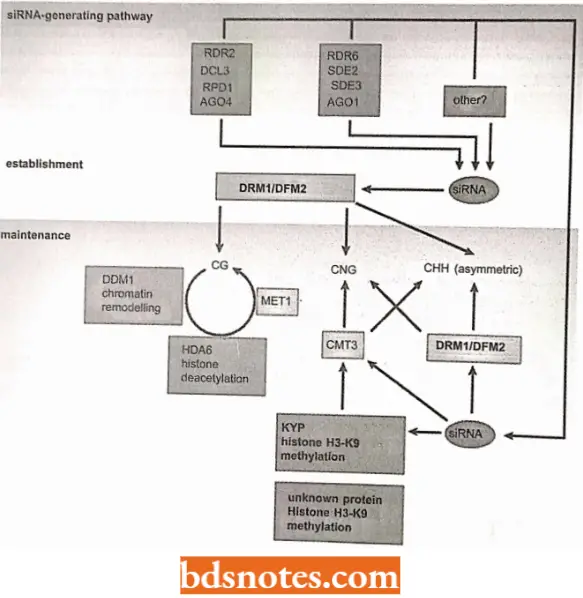
Methylation of DNA: The importance of methylation in DNA-protein interactions is well known. Molecular biologists have shown that a particular DNA sequence could be protected from restriction endonucleases if it was methylated.
- A small percentage of cytosine residues are methylated in many eukaryotic organisms, mainly in CpG sequences; 80% of the cytosines in CpG sequences in human DNA are methylated. (Note. Often when we refer to a sequence of two bases on the same strand of DNA, we put a “p” between them -CpG to indicate that they are on the same strand connected by a phosphodiester bond and not on two different strands as a hydrogen-bonded base pair).
The Degree Of Methylation Of DNA Is Related To The Silencing Of Genes: Genes that are dormant at one stage of development but active in another, are usually less methylated when active and more fully methylated when inactive. For example, adenovirus, a cancer-causing virus, has been observed in many eukaryotic cell lines.
- In most lines in which the adenovirus DNA has integrated into the host chromosome, late viral genes are turned off. These genes are highly methylated at their CCGC or GCGC sites.
- In addition, chemicals that prevent methylation frequently activate previously dormant genes.
- For example, a chemical compound, called 5-azacytidine, inhibits methylation; X chromosomal genes, which are normally deactivated, can be reactivated by treatment with 5-azacytidine.
- There are numerous other examples of the activation of genes after treatment with this chemical.
- The activated genes lack methylated cytosines that were previously methylated. Finally, the possibility exists that DNA methylation can affect chromatin structure.

- Recent work has also indicated that the methylation itself may not prevent transcription, but milter may be a signal for transcriptional inactivity.
- In the talc cress plant, Arabidopsis thaliana a protein named Mom (for Morpheus molecule) has been discovered that, when mutated, results in cones that have heavy methylation levels but are actively transcribed.
- Thus, the methylation level can be separated from the transcriptional activity of genes, although the two usually occur together.
- Ambidopsis is proving to be a good model in the study of the role of methylation in transcriptional activation because of other common model organisms, namely fruit flies, yeast, and the nematode.
- Caenorhabditis elegans. do not have methylation of their DNA.
- Further interest has been generated in the role of methylation in regulating gene expression by the discovery of Z DNA, and the fact that Z DNA can be stabilized by methylation.
- This observation has led to a model of transcriptional regulation based on alternative DNA structures. Sequences (such as CpG repetitions) that could exist as Z DNA exist as B DNA when being transcribed.
- If the gene is to be silenced (turned off), the CpG sequences are converted to stable Z DNA by methylation, which then blocks transcription.
- This feasibility has gained some interest because of the recent discovery of an enzyme, double-stranded RNA adenosine deaminase(ADARl), that binds to Z DNA sequences.
“Why are transcription mechanisms often misunderstood in practice? Questions answered”
Signal Transduction: Let us see, how specific transcription activation factors appear at specific times. During the development of animals, control of gene expression requires that genes be expressed at specific times and under specific circumstances.
- If transcription is usually controlled by specific transcription factors, what determines the appearance of these factors at the appropriate times and places? For both of these requirements, there exists one common mechanism, called signal transduction pathway, in which signals pass from the external environment through the cytoplasm, into the nucleus.
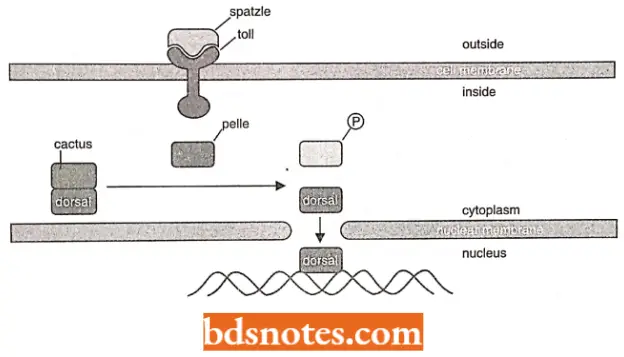
- For example, in a signal transduction pathway involved in the development of the fruit fly, the Toll protein spans the plasma membrane.
- It acts as a receptor for the Spatzle protein, which when detected, causes a change in the cytoplasmic end of Toll protein, activating it.
- Activated Toll protein activates Pelle, a protein kinase that phosphorylates the Cactus protein, causing it to dissociate from Dorsal protein.
Once the Dorsal protein dissociates from the Cactus protein (which acts to repress the Dorsal protein), the Dorsal protein becomes an active specific transcription factor that can cross the nuclear membrane and activate its target genes.
- We thus see that the Spatzle protein attaching to its receptor protein (i.e., Toll protein) on the cell surface results in the activation of the target gene of the Dorsal protein in the nucleus.
- These pathways can become very complex, with many protein elements. More elements mean more sensitive control of various processes, often requiring that several conditions be met before a gene is activated.
- In addition, these pathways are usually conserved by evolution.
- A similar pathway, though more complex, occurs in mammals in which the target gene is interleukin-1, a protein in the immune system that induces fever.
- In the mammalian protein, the signal protein is called Toll-like receptor-4 and the specific transcription factor is called NF-kB.
Transcription elongation produces superhelical tension of DNA: Once it has initiated transcription, RNA polymerase does not proceed smoothly along a DNA molecule, rather it moves jerkily, pausing at some sequences and rapidly transcribing through others.
- RNA polymerases involved in RNA elongation both bacterial and eukaryotic, are associated with a series of elongation factors, proteins that decrease the likelihood that RNA polymerase will dissociate before it reaches the end of a gene.
- These factors typically associate with RNA polymerase shortly after initiation has occurred and help polymerases to move through the wide variety of different DNA sequences that are found in genes.
- Eukaryotic RNA polymerases must also cope with chromatin structure as they move along a DNA template.
- Experiments have shown that bacterial RNA polymerases, which never encounter nucleosomes in vivo, can nonetheless transcribe through them in vitro, suggesting that a nucleosome easily traverses.
- However, eukaryotic polymerases have to move through forms of chromatin that are more compact than a simple nucleosome.
- It therefore seems likely that they transcribe with the aid of chromatin remodeling complexes.
- These complexes may move with the RNA polymerases or may simply seek out and rescue the occasionally stalled RNA polymerase.
In addition, some elongation factors associated with eukaryotic RNA polymerase facilitate transcription through nucleosomes without requiring additional energy.
- It is not yet understood how this is accomplished, but these proteins may help to dislodge parts of the nucleosome core as the RNA polymerase transcribes the DNA of a nucleosome.
- Elongating RNA polymerases also have to phase out some other types of barrier, both bacterial and eukaryotic. To discuss this issue, we need first to consider a refined property inherent in the DNA double helix called DNA supercoiling.
- DNA supercoiling represents a conformation that DNA will adopt in response to superhelical tension; conversely, creating various loops or coils in the helix can create such tension.
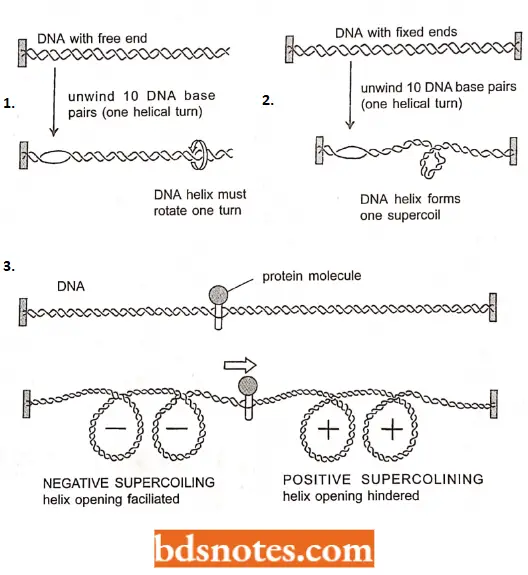
- A simple way of visualizing the topological constraints that cause DNA supercoiling is illustrated. There are approximately 10 nucleotide pairs for every helical turn in a DNA double helix.
- In a gene a helix whose two ends are fixed concerning each other (as they are in a DNA circle, such as a bacterial chromosome, or a tightly clamped loop, as thought to exist in eukaryotic chromosomes).
- In this case, one large DNA supercoil will form to compensate for each 10 nucleotide pairs that are opened (unwound).
- The formation of this super coil is energetically favorable because it restores a normal helical twist to the base-paired regions that remain which otherwise need to be overwound because of the fixed ends.

- Superhelical tension is also created as RNA polymerase moves along a stretch of DNA that is anchored at both ends.
- As long as the polymerase is not free to rotate rapidly (and such rotation is unlikely given the size of DNA polymerases and their attached transcripts), a moving polymerase generates positive superhelical tension in the DNA in front of it and negative helical tension behind it.
- For eukaryotes, this situation is thought to provide a bonus: the positive superhelical tension ahead of the polymerase makes the DNA helix more difficult to open but this tension should facilitate the unwrapping of DNA in nucleosomes, as the release of DNA from the histone core helps to relax positive superhelical tension.
- Any protein that propels itself along a DNA strand of a double helix tends to generate superhelical Chromatin remodeling complexes.
“Cost of ignoring transcription principles vs benefits of systematic approaches: Q&A”
- In eukaryotes, the DNA topoisomerase enzyme rapidly removes this superhelical tension.
- But, in bacteria, a specialized topoisomerase called DNA gyrase uses the energy of ATP hydrolysis to pump super oils continuously into the DNA, thereby maintaining the DNA under constant tension.
- These are negative supercoils, having the opposite handedness from the positive supercoils that form when a region of the DNA helix opens.
- These negative supercoils are removed from bacterial DNA whenever a region of helix opens, reducing the superhelical tension.
- DNA gyrase therefore makes the opening of the DNA helix energetically favorable compared with the helix opening in DNA that is not supercoiled.
- For this reason, it usually facilitates those genetic processes in bacteria, including the initiation of transcription by bacterial RNA polymerase, that require helix opening.
Transcription Questions And Answers
Question 1. Describe the differences, if any, between the chemical reactions catalyzed by DNA polymerase and RNA polymerase.
Answer:
The reactions are identical, however, substrates are different, i.e., DNA polymerase joins deoxynucleotides and RNA polymerase joins ribonucleotides.
Question 2. What is a transcription unit? Is it the same thing as a gene?
Answer:
A transcription unit is a section of DNA extending from a promoter to an RNA polymerase termination site. It is usually not a gene but typically includes many genes.
Question 3. In what way, relating to polycistronic mRNA, do eukaryotic and prokaryotic protein synthesis differ?
Answer: In eukaryotes all mRNA is monocistronic.
Question 4. What is meant by the terms “upstream” and “downstream”?
Answer:
Upstream and downstream usually refer to regions in the 5 and 3′ directions respectively, from a particular site that is being discussed.
Question 5. An RNA molecule is isolated having a 3′-OH terminus and a 5-P terminus. What information does this fact provide?
Answer: Since the triphosphate of the primary transcript is absent, the molecule has been processed
Question 6. What is a consensus sequence? What is a conserved sequence?
Answer:
A Consensus sequence is made up of the nucleotides that appear in a significant proportion of cases when similar sequences are aligned. A conserved sequence consists of nucleotides found in all cases when similar sequences are aligned. For example, the Pribnow box is the consensus sequence TATAAT.
Question 7. Why do you think that most promoter regions are A-T rich?
Answer:
A double helix of DNA must unwind for transcription to occur. A-T pairs, because they have only two H-bonds, are more easily disrupted than G-C pairs.
Question 8. What is the difference between a G-70 and a G-32?
Answer:
The superscripts of the sigma factors refer to their molecular weights (for example., cr-70 is 70,000 daltons). Different sigma factors usually recognize different prokaryotic promoters.
Question 9. What is footprinting? How did it help to define promoter?
Answer:
- Footprinting is a technique in which DNA in contact with a protein is exposed to nucleases; only DNA protected by the protein is undigested.
- Promoters could be isolated by protection with RNA polymerase in the absence of ribonucleotides- the polymerase will not move- and then sequenced.
Question 10. Can one nucleotide be a conserved sequence?
Answer:
- Conserved sequences are invariant sequences of DNA or RNA recognizable to either a protein or a complementary sequence of DNA or RNA. However, in group 2 introns, an adenine is needed near the 3′ end of the intron for lariat formation.
- Thus, this single nucleotide, given its relative position in the intron and possible surrounding bases, is a conserved sequence of one.
“Success rate of interventions using modern transcription techniques: FAQ”
Question 11. What is a stem-loop or hair-pin structure? An inverted repeat? A tandem repeat? Draw a section of a DNA double helix with an inverted repeat of seven base pairs.
Answer:
A stem-loop (or hairpin) structure can form when a single strand of DNA or RNA has a double helical section. An inverted repeat is a sequence read outward on both strands of a double helix from a central point. A tandem repeat is a segment of nucleic and repeated consecutively; that is, the same sequence repeats in the same directions on the same strand:
- 5′- TCCGGTCCGGTCCGG- 3′
- 3′- AGGCCAGGCCAGGCC- 5′
A DNA sequence with a seven-base inverted repeat is
- 5′- ATTACCGCGGTAAT-3′
- 3′- TAATGGCGCCATTA-5′
Transcription Multiple Choice Questions And Answers
Question 1. Transcription is the transfer of genetic information from
- Chromosome’to cytoplasm
- TRNAtomRNA
- DNAtomRNA
- mRNA to rRNA
Answer: 3. DNAtomRNA
Question 2. Nongenetic RNA is of
- One type
- Two types
- Three types
- Nonfunctional type
Answer: 3. Three types
Question 3. Regarding transcription which one of these is false?
- Uses RNA polymerase
- One of the mother strands of DNA is the template
- Is bidirectional
- Reverse transcription uses RNA-dependent DNA polymerase
Answer: 3. Is bidirectional
Question 4. The enzyme required for transcription is
- DNA polymerase
- RNA polymerase
- Endonuclease
- All the above
Answer: 2. RNA polymerase
Question 5. Pribnow box contributes to
- Protein synthesis
- ATP synthesis
- RNA synthesis
- None of the above
Answer: 3. RNA synthesis
Question 6. The process involved in the RNA formation on DNA template is
- Translation
- Transduction
- Transcription
- Transformation
Answer: 3. Transcription
Question 7. Transcription takes place in
- Matrix
- Nucleus
- Cytosol
- Cytoplasm
Answer: 2. Nucleus
Question 8. Essential components of eukaryotic cistron are
- Intron
- Exons
- Operons
- Operator and regulator genes
Answer: 4. Operator and regulator genes
Question 9. After initiation of transcription with core enzyme RNA polymerase the sigma factor is
- Functionlcss
- Released to take part again
- Used during the closing of the chain
- Retained and it performs a special function
Answer: 2. Released to take part again
Question 10. During transcription, the DNA site at which RNA polymerase binds is called
- Promoter
- Regulator
- Receptor
- Enhancer
Answer: 1. Promoter
Leave a Reply