Regulation Of Gene Action In Eukaryotes
The different types of cells in the body of a multicellular organism differ in structure and function, but their genes are identical since all cells are ultimately derived from the zygote by cleavage (mitosis). The problem therefore is: how do cells with identical genetic complements differ so much in stature and function? The answer is that not all genes are active at one time.
- As development proceeds certain genes become active while others become inactive; i.e., the genes are “switched on” and “switched off” at different times. This process is called differential gene action. When genes are active they direct the formation of enzymes that affect certain phenotypic traits.
- Further, though a cell has the genes to produce hundreds of enzymes, only the enzymes required at a particular time are produced. Such a control mechanism ensures that the cell is not Hooded with unnecessary enzymes.
- In eukaryotes, genes are active only when their products are required by the cell. The rest of the time they are “switched off. For example, in mouse liver cells, only about 3% of the genes are active (i.e., transcribed RNA), and in brain cells, about 9% of the genes are turned on.
- In eukaryotes, the following two kinds of controls or regulations of gene expression occur:
- Short-term or reversible regulation corresponds to the kind of regulation we studied in bacteria and it represents a cell’s response to fluctuations in the environment, especially, it involves changes in activities of concentrations of enzymes as particular substrates and or hormone levels rise and fall.
- The changes a cell experiences during a cell cycle, particularly the fluctuations in rates of DNA, RNA, and protein synthesis that regularly occur concerning the time of mitosis can also be placed in this category.
- Long-term or irreversible regulation includes the phenomena associated with determination, differentiation, or more generally development: it is involved in the numerous steps by which a fertilized egg becomes an organism of, perhaps, trillions of cells with diverse and ultimately quite permanent roles to play in the maintenance of the whole. Short-term regulations also occur in developing and differentiating eukaryotic cells side by side with long-term regulation.
Both of these types of regulations of gene activities in eukaryotes, now, are considered to occur at the following levels involving diverse mechanisms:
- Regulation at the level of DNA;
- Regulation at the level of transcription;
- Regulation at the level of translation; and
- Regulation at the level of post-translation.
“Understanding eukaryotic gene regulation through FAQs: Q&A explained”
Regulation Of Gene Action At The Level Of Genome
In eukaryotic cells, it seems that certain classes of genes are transcribed more or less continuously, and only in extreme situations their activities are repressed. For example, genes coding for larger ribosomal RNA (28S or 18S rRNA)) or transfer RNA (tRNA) are present as multiple copies forming a simple multigene family.
- These genes are transcribed uniquely by RNA polymerase I for the larger ribosomal RNA or by RNA polymerase 3 for tRNA and 5S RNA. Although the products of some of these genes, the ribosomes, are used continuously in all cells, it does not confirm that all of these multiple copies are continuously transcribed at maximum rate.
- Electron micrographs of spread chromatin from nucleoli often show that some of the repetitious rRNA genes are inactive. It is also true that in the nucleated erythrocytes of lower vertebrates such as Xenopus, all genes may be turned off (Maclean et al., 1972), including those for ribosomal RNA and tRNA.
- Therefore, it is clear that mechanisms do exist for inactivating sequences even those regarded to be constitutive in normal cells. Some of the clearest demonstrations that some specific genes are at least available for transcription in different kinds of differentiated cells are provided by Drosophila and other organisms.
- For example, the pattern of bands and interbands of polytene chromosomes of Drosophila does not vary between different larval tissues, yet it is now concluded that the interband regions probably represent housekeeping genes that code for essential proteins, that they are expressed in every cell, and that they are retained in a state of permanent decondensation (Bautz and Kabisch, 1983).
- Thus, ‘housekeeping’ genes may be ‘left on’ for much of the life of the cell when transcription of even the most essential housekeeping genes ceases (i.e., during mitosis).
- Further, when we consider the case of the “cell-specific” genes, luxury genes or smart genes, which code for the products only found in specialized tissues, it becomes immediately clear that differential expression is the rule.
- Whether expression of a gene is measured at the level of the messenger RNA or the protein, genes coding for products such as globin, crystallin, fibroin, ovalbumin, casein, and immunoglobulin give every indication of complete repression in all but the specialized tissue characterized by their presence.
- Thus, at the level of the genome (i.e., DNA), the following five modes of regulation are operative:
The Situation Of The Total Genetic Shutdown
- During the mitotic phase of the cell cycle, chromatin is highly condensed to form chromosomes, and the transcriptional activity of all genes is suspended.
- During the meiotic division of germ cells, a somewhat similar situation to (a) is evident, although in some rare cases, such as lampbrush chromosomes of meiotic diplotene in vertebrates (Vlad, 1983), transcription proceeds very actively.
- The nucleus of mature nucleated erythrocytes of amphibians is transcriptionally inactive. Chromatin in these cells is highly condensed but not organized into discrete chromosomes (Chegini et al., 1981).
- However, transcription can be partially reactivated in these nuclei by transferring them into new cytoplasm or exposing them in vitro to altered environmental conditions.
- In mammalian females, one of the two X chromosomes present in somatic cells undergoes condensation in early embryonic stages to become heterochromatic sex chromatin or Barr body (Dosage compensation).
- A variety of experiments indicate that most, though not all, genes of the condensed X chromosome are turned off. In developing oocytes, as opposed to somatic cells, Barr bodies are not present, the activities of both X chromosomes being required for normal oogenesis.
- In somatic cells of normal XY males, genes of the single X chromosome remain active and Barr bodies are not found. However, in germ cells, the X chromosome is inactivated before spermatogenesis, otherwise, it may prevent sperm maturation and lead to sterility*. In one extreme case, that of the creeping vole.
- Micivfus Oregon, the X chromosome is eliminated from the germ cells of males by a special process of nondisjunction.
- Sperm cells contain a complete genetic endowment but no transcription occurs until the sperm nucleus is activated within the egg cytoplasm.
- Complete suspension of transcriptional activity* is also known in the following cases: cells of some plant seeds; cells within diapausing Anemia gastrulae; cells within inactive organisms such as desiccated Tardigrada nuclei within bacterial and fungal spores; and nuclei within desiccated amoeba cells, as in the slime mold Dictyostelium.

“Importance of studying eukaryotic gene regulation for biology students: Questions explained”
Evidence For Constitutive Expression Of Some Genes
- If the interbands of Drosophila polytene chromosomes are correctly interpreted as being loci for “housekeeping” genes, then the evidence is that such chromatin is permanently decondensed and is transcribed at a low but constant rate (Semehin et al.. 1979).
- Electron microscopy of spread films of DNA extracted from nucleoli of laevis oocytes reveals tandemly arranged sequences coding for the 45S precursor of ribosomal RNA, each gene adorned with a Christmas-tree arrangement of RNA in the process of synthesis (i.e., transcription).
- There is a constant and universal requirement for the products of certain genes in all cells and at all times. These include products such as the four kinds of rRNA-28S, 18S, 5.8s, and 5S; tRNA of 20 basic types, and a few hundred proteins such as histones, ubiquitin and lactate dehydrogenase, RNA polymerase, and the like.
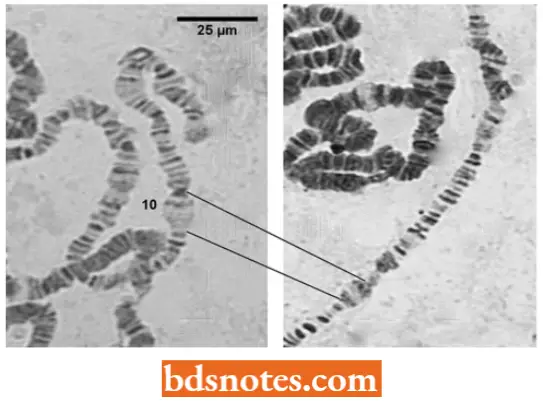
Many Genes Are Expressed Only In Certain Tissues
- Xenopus provides a good example of the regulation of 5S genes. Xenopus borealis possesses 19,000 copies of the oocyte-specific 5S rRNA genes, and these genes are active only in the oocyte and in no other cell.
- The enzyme lactate dehydrogenase (LDH) is coded by a small family of genes, each gene determining the structure of a subunit. Subunits A and B are expressed in almost all mammalian cells, but one of the genes in the family, coding for subunit C, is active only in spermatocytes within the developing testes.
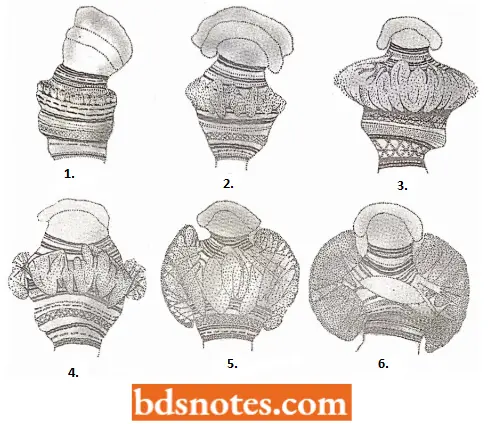
- The puffing (i.e., chromatin decondensation) of restricted segments of the polytene chromosomes of Drosophila, Chironomus, etc., provides visible evidence of the activity of genes coding for cell-specific products.
- Certain puffs, known as heat-shock puffs, can be induced to appear specifically when salivary glands are exposed to heat shock either in vivo or in vitro. The correlation between such chromatin decondensation and transcription activity is proved by autoradiography using tritiated precursors of RNA.
- Some DNA is never transcribed in any cell. Analysis of various types of DNA sequences existing in eukaryotic cells reveals that some DNA is comprised of tandemly repeated short sequences that are concentrated in heterochromatin such as centromeres of chromosomes and the Y chromosome. Current evidence indicates that much of this DNA is never transcribed in any cell.
- Some spacer sequences occur between genes, for example, between multiple copies of genes for ribosomal RNA. Such spacer sequences are often taken to be untranscribed, but now it is found that some of the spacer DNA may transcribe nuclear RNA molecules.
The very large size of the genomes of the higher eukaryotes certainly indicates that much of the DNA is redundant (=repetitious) and probably not utilized as a coding or regulatory sequence.
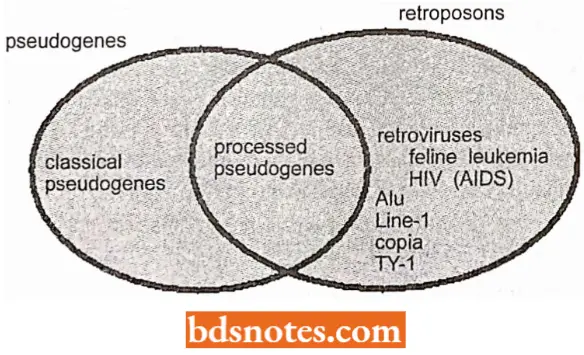
“Common challenges in understanding eukaryotic gene regulation effectively: FAQs provided”
- The pseudogenes found in many gene families are often presumed to have arisen as cDNA DNA) copies of reverse transcribed messages. They lack introns and contain many stop signals so that RNA polymerase molecules fail to move very far along them.
- Some DNA is spliced to cause gene rearrangement. Such a mechanism occurs during the expression of immunoglobulin (Ig) genes.
Regulation Of Gene Action At The Level Of Transcription
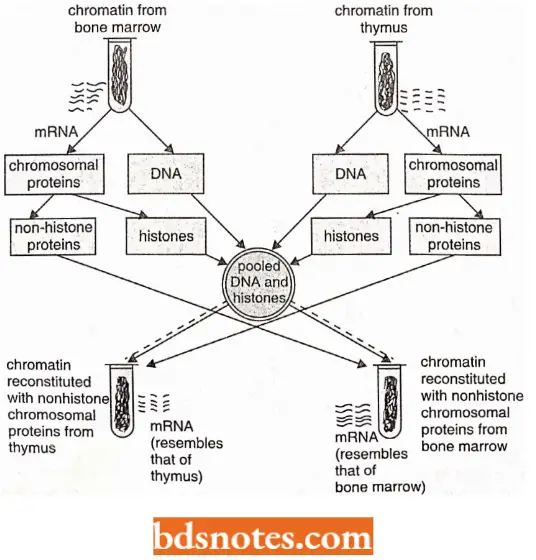
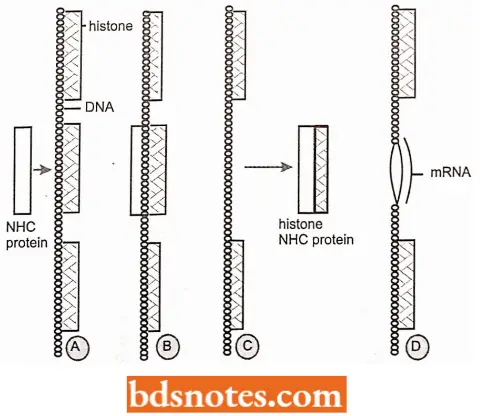
Chromatin reconstitution experiments. Chromatin has main components—DNA, histones, and non-histones. While it has been known since the early 1960s that histones may be involved in repressing gene activity, the specific regulation by non-histones was shown only during the 1970s.
- Gilmour and Paul (1973) performed a chromatin reconstitution experiment to demonstrate the positive role of non-histones in the regulation of gene activity.
- They isolated the chromatin from different tissues separately and then dissociated it into DNA, histones, and non-histones.
- This is followed by chromatin reconstitution using either the three components derived from the chromatin of the same tissue or by combining the non-histones of one tissue with the DNA and histones of another tissue.
- From such experiments, it was demonstrated that the mRNA which is synthesized in vitro from reconstituted chromatin, mainly depended on the source of non-histone proteins.
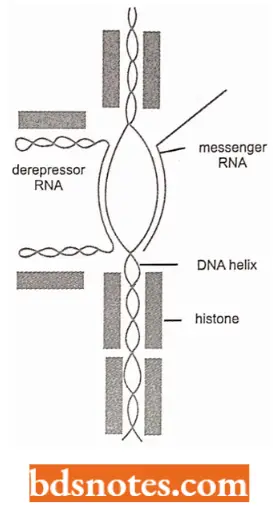
- Freusters(1965) model of gene-specific depressor UNA. Histones stabilize the DNA double helix, by interacting with the negative phosphate groups of DNA. This prevents the separation of strands. and consequently transcription. The histones, thus, net as a general repressor of protein synthesis.
- Histones are displaced from DNA by nuclear polyions which form complexes with histones. This permits the separation of the DNA strands in a random manner.
- Gene-specific derepressor RNA hybridizes with the non-transcribing DNA strand at a particular locus and stabilizes the loop in the open position.
“Why is early learning of gene regulation critical for molecular biology? Answered”
- This frees the other DNA strand and permits it to transcribe mRNA. Stein’s model. According to G.S. Stein, J.S. Stein and Kleinsmith (1975). individual NHC (nonhistone chromosomal) proteins recognize certain sites on DNA and bind to it there.
- They then pull off the histone repressor from the site, thus, derepressing DNA. Thus, NHC proteins act by derepressing.
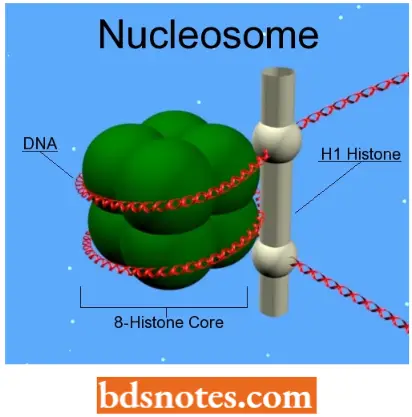
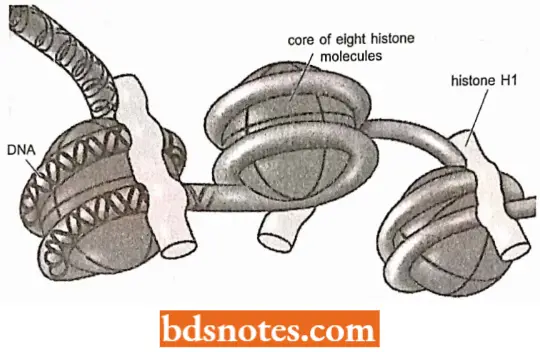
Change In Chromatin Conformation: Nucleosomes continue to be present on most transcriptionally active DNA sequences, but they are probably reduced in number.
- Thus, although evidence from some laboratories suggests that the ribosomal genes of the amphibian nucleolus lack nucleosomes, active gene loci in Drosophila give a positive reaction to antibodies against H3 and H4, indicating that at least these subunits of the nucleosome persist on such DNA.
- In fact, in some active genes, the nucleosomes are displaced or “phased” in these regions(Samal et. al., 1981).
Modification of DNA sequences: DNA methylation. The genomic DNA of higher eukaryotes is modified following replication so that a large proportion of the cytosine (C) residues are present as 5-methylcytosine(5mC).
- However, such methylation has not been detected in the DNA of lower eukaryotes such as yeast and Dictyostelium, nor Drosophila.
- The percentage of methylated C residues inDNA relative to unmethylated C residues is highly variable from less than 1 percent in some insects to over 50 percent in some higher plants and vertebrates.
- A much greater correlation exists between the methylation or under-methylation of sequences in the vicinity of gene promoters.
- For example, the DNA of sperm is highly methylated, as in the DNA of the oocyte-specific 5S rRNA genes in adult tissues, whereas the sites around the coding regions of genes such as adult globin, ovalbumin, and immunoglobulin are under-methylated in tissues in which they are expressed but are largely methylated in other cells in which they are not expressed.
- There is evidence that the cytosine methylation in DNA alters the structure of the double helix fundamentally and favors the transition from B-form to Z-form DNA (Bele and Felsenfed, 1981). It is possible that B↔Z transition is itself involved in gene regulation and this may be how DNA methylation has its effects on transcription.
Modifications Of Histones: The histone component of chromatin is subject to three different post-synthetic modifications that have either direct or indirect effects on eukaryotic gene regulation:
- Histone methylation affects only histones H3 and H4 and involves the irreversible methylation of a few lysine residues which alters the hydrophobic nature of the side chain of these histones.
- Histone phosphorylation involves phosphorylation that affects serines and threonines, changing them from a state of neutral charge to one of negative charge, and is a reversible reaction.
- The state of phosphorylation of HI protein varies through the eukaryotic cell cycle, and after HI phosphorylation, chromatin becomes much more strongly condensed, as it does in mitotic chromosomes.
- activation of the histone kinase enzyme that is responsible for H 1 phosphorylation may be the first step in the chain of events that leads to eventual chromatin condensation before mitotic cell division.
“Factors influencing success with eukaryotic gene regulation knowledge: Q&A”
- Histone acetylation is of two types. The first is the irreversible acetylation of the amino-terminal serines of histones H1, H2A, and H4.
- These modifications seem to be associated with histone synthesis. The second is the reversible acetylation of lysine residues in the aminoterminal regions of histones H2A, H2B, H3, and H4.
- Acetylation converts the normally basic lysine side chain to a neutral acetyl-lysine, and, thus, reduces the net basic charge of the aminoterminal ends of the affected histones.
- Both H3 and H4 can have up to four lysines in the acetyl form, and there is a strong correlation between this type of histone acetylation, especially deacetylation of all available lysines, and transcriptionally active chromatin.
- The acetylation of the core histone lysines would tend to loosen the nucleosomal structure which brings about the transition from a silent condensed gene to a transcriptionally active and extended one.
Transcriptional Regulation By Protein A24: The A24 is an unusual hybrid protein, being a complex of histone H2A and the non-basic protein ubiquitin.
- The ubiquitin is covalently bound via the side chain amino group of lysine 19 of the histone.
- Some 10 percent of H2A molecules are in the form of A24 and these specialized histones seem to be confined to interphase chromatin, disappearing as the chromosomes condense.
- A24 is found highly abundant in the chromatin of active genes.
Gene Regulatory Molecules: Transcription of the eukaryotic genome is believed to be regulated by a variety of specific gene regulatory molecules which are produced by specific regulatory genes or by cytoplasm/cell surface. Examples of such gene regulatory molecules are the following:
- RNA polymerases. These enzymes are necessary for transcription and if they are short in supply, they tend to affect it. For example, there is a possible competition for type 2 polymerase (which is meant for hnRNA and mRNA) by the various promoter sequences that lie upstream of the protein-coding sequence.
- Endonucleases. These enzymes are likely to affect the transcription, especially in vitro cell-free systems, by introducing nicks into DNA that may serve as initiation sites for polymerases.
- Topoisomerases, helicases, and other DNA helix-destabilizing proteins. Various proteins are known that alter the three-dimensional structure of DNA and render it more available for processing which may affect transcription.
- DNA Methylase. The enzyme is likely to make DNA less available for transcription, and factors that antagonize methylation would enhance transcription.
- Histone acetylase and deacetylases. Such enzymes influence the rate of transcription by modulating acetylation.
- Factors such as ATP. Such molecules may influence the transcription rate by changing the available energy.
- Ions and small molecules. Many ions such as those of calcium, magnesium, and manganese directly affect chromatin conformation, which modulates gene activity.
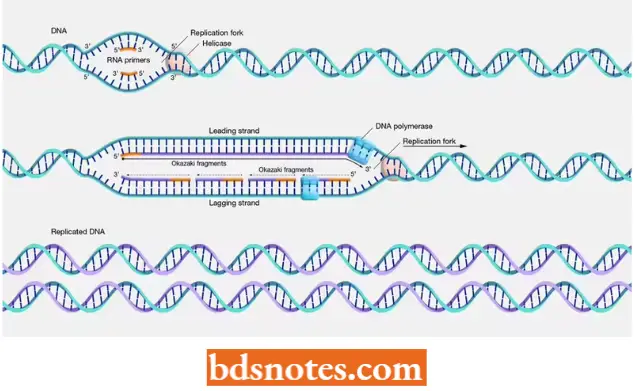
“Steps to explain eukaryotic gene regulation: Transcriptional vs post-transcriptional control: Q&A guide”
Britten-Davidson model or gene-battery model of transcription regulation. This model of eukaryotic gene regulation at the level of transcription was proposed by Britten and Davidson in 1969 and later on, elaborated by them in 1973. The gene-battery model assumes the presence of four classes of sequences:
- Producer gene which is comparable to a structural gene of prokaryotic operon;
- The receptor site is located adjacent to each producer gene and is comparable to the operator gene of the prokaryotic operon;
- The integrator gene is comparable to the regulator gene and is responsible for the synthesis of an activator RNA that may or may not give rise to proteins before it activates the receptor site;
- The sensor site regulates the activity of the integrator gene, which can be transcribed only when the sensor site is activated. The sensor sites are recognized by agents which, like hormones and proteins, change the pattern of gene expression.
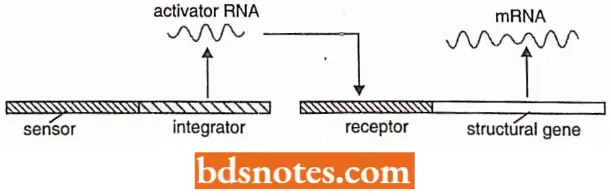
- For instance, a hormone protein complex or a transcription factor may bind to a sensor site and cause the transcription of the integrator.
- In this model, the genes (producer gene and integrator gene) are those sequences that are involved in RNA synthesis, while the receptor and sensor sites help only in recognition without taking part in RNA synthesis. Lastly, in Britten-Davidson’s model, a set of structural genes controlled by one sensor site is called the battery.
Post-Transcriptional Regulation
Steps that come between transcription and translation are described as post-transcriptional and are the following:
- Some RNA is capped and tailed. The precise functions of capping and tailing of mRNA are not known, but they seem to serve to identify a message or potential message, and tailing may also help in the final export of this message from the nucleus.
- The mRNA is capped at its 5′ end by modified guanosine residue. Messenger RNAs that have unmodified caps are not translated. For example, stored mRNA in the oocyte of the tobacco hornworm moth has the guanosine added to its 5′ end, but the G is not modified.
- Hence, this stored mRNA is not translated. However, after fertilization, the cap is modified, allowing the mRNA to be translated to produce proteins needed for early embryogenesis (see Purves et al., 1998).
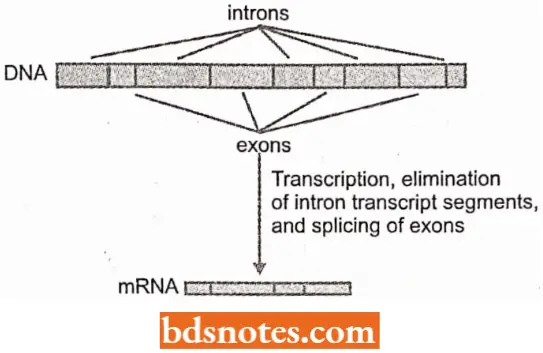
- RNA is processed to remove intron sequences. Introns removal and splicing together of the remaining exons during the processing of hnRNA must be absolutely precise. This is in part engineered by a distinct group of nuclear particles(snRNPs containing U1, U2, U3, U4, U5, and U6 snRNAs).
- For example, differential splicing is used in different lymphocyte cells to produce different proteins from the same hnRNA molecule. As originally discovered by Early et al., (1980), the two types of mRNA molecules are produced by part of an intron being omitted from one of the mRNAs but included in the exon splice used to produce the other mRNA.
- This allows the production of two distinct proteins both immunoglobulins (Ig), but one with a long strand of hydrophobic amino acids at its carboxyl terminus, and the other with only a short length of relatively hydrophilic amino acids. The Ig molecule with the long hydrophobic peptide is secreted from the cell.
- This change in splicing takes place within the life of a single lymphocyte cell and clearly explains the following observation. Immature lymphocytes retain antibodies and simply insert the Ig molecules into their plasma membranes, whereas following stimulation with antigen, the same lymphocyte becomes secretory’, releasing antibody molecules into circulation.
- Most RNA is never exported from the nucleus. About 5 percent of total transcribed RNA never leaves the nucleus. This is explained partly by the removal of intron RNA and also by many RNA molecules that break up within the nucleus.
- The significance of this process is not clear, but some clues about the identification of RNA for export are coming to light.
- Although not all genes contain introns, most do, and it seems that the presence of some of these introns is essential for RNA export. In other words, introns are used as a means of identifying or ticketing the molecules that are to be passed out of the nucleus (see Maclean and Hall, 1987).
- Message degradation rates are significant. The rate at which eukaryotic mRNA is degraded in the cytoplasm is highly variable. This implies that differential message breakdown is an important method of regulating not only the rate of gene expression but also the lag between transcriptional shutdown and the cessation of specific translation.
- For example, the survival of histone mRNA during the cell cycle explains this fact very clearly. New histone is required in massive amounts immediately at the start of the S period of DNA synthesis to provide the new DNA with nucleosomes.
- Recently, it was discovered that the restricted availability of histone messages is not achieved as a result of transcriptional control alone but by differential breakdown rates for histone messages.
Tubulin is a protein that polymerizes to form microtubules, a component of the cytoskeleton. When a large pool of free tubulin is available in the cytoplasm, there is no particular need for the cell to make more of it.
- Under these conditions, some tubulin molecules bind to tubulin mRNA, and this binding makes such mRNA molecules especially susceptible to breakdown, and less tubulin is made.
- Other examples illustrate the same mechanism- that the less time an mRNA stays in the cytoplasm, the less it can be translated into protein (see Purves et al., 1998).
“Role of transcription factors in eukaryotic gene regulation: Questions answered”
Translational Control
In bacteria, most mRNA molecules are translated about the same number of times with only fairly small variation from gene to gene. In eukaryotes, translational regulation occurs in which a mRNA molecule is not translated at all until a signal is received. Translational control may involve the following mechanisms:
Extension Of Lifetime Of The mRNA: An important example of translational regulation is that of informosomes or masked mRNA.
- Unfertilized eggs are biologically static, but shortly after fertilization, many new proteins must be synthesized, for example, the proteins of the mitotic apparatus, the cell membranes, histones for nucleosome formation as well as others.
- Unfertilized sea urchin eggs store large quantities of mRNA for many months in the form of mRNA-protein particles(ÿmasked mRNA) made during the formation of the egg.
- This mRNA is translationally inactive, but within minutes after fertilization, translation of these molecules begins. Here, the timing of translation is regulated; the mechanism for stabilizing the mRNA, for protecting it against RNases, and for activation are still unknown.
Regulation Of Rate Of Protein Synthesis: This type of regulation also occurs in mature unfertilized eggs. These cells need to maintain themselves but do not have to grow or undergo a change of state. Thus, the rate of protein synthesis in eggs is generally low.
- This is not due to an inadequate supply of mRNA but to a limitation of an as-yet-undefined element, called the recruitment factor which interferes with the formation of the ribosome-mRNA complex.
- A good example of translational control is the extension of the lifetime of silk fibroin mRNA in the silkworm Bombyx mori. During cocoon formation, the silk gland of the silkworm predominantly synthesizes a single type of protein, silk fibroin.
- Since the worm takes several days to construct its cocoon, it is the total amount and not the rate of fibroin synthesis that must be great; the silkworm achieves this by synthesizing a fibroin mRNA molecule that is very long-lived.
- Transcription of the fibroin gene is initiated at a strong promoter by an unknown signal and about 104 fibroin mRNA molecules are made in several days(such a synthesis forms an example of transcriptional regulation).
A typical eukaryotic mRNA molecule has a lifetime of about three hours before it is degraded. However, the fibroin mRNA survives for several days during which each mRNA is translated repeatedly to yield 105 fibroin molecules.
- Thus, each gene is responsible for the synthesis of 109 protein molecules in four days. Altogether the silk gland makes 300μg or 1015 molecules of fibroin during this period.
- If the lifetime of mRNA were extended, either 25 times as many genes would be needed or synthesis of the required fibroin would take about 100 days.
- Further, free iron ions (Fe2+) within a mammalian cell are bounded by a storage protein, ferritin. When iron is in excess, ferritin synthesis rises dramatically. Yet the amount of ferritin mRNA remains constant. This increase in ferritin synthesis is due to an increased rate of mRNA translation.
- When the iron level in the cell is low, a translational repressor protein binds to ferritin mRNA and prevents its translation by blocking its attachment to the ribosome.
- When the iron level rises, the excess iron binds to the repressor and alters the three-dimensional structure of the translational repressor, causing it to detach from the mRNA and translation proceeds (Purves et al, 1998).
Post-Translational Modification Of Proteins To Make Them Active Ones
Some proteins are altered after synthesis, usually by partial degradation or trimming, as, by the enzymatic removal of the central section of the proinsulin molecule to yield the active protein, insulin.
- For their activity, many proteins also depend on being complexed into compound proteins together with other subunits, either the same or different.
- Such posttranslational control mechanisms do play a significant role in determining the activities of differential cells.
- For example, hemoglobin production is highly dependent on the availability of haem to complex with globin protein subunits which may be deficient in cases of iron-dependent anemia.
Hormonal Control Of Gene Expression
In higher plants and animals, intercellular communication is a very important phenomenon. Signals originating in various glands and or secretory cells somehow stimulate target tissue or target cells to undergo dramatic changes in their metabolic patterns.
- These changes frequently include altered patterns of differentiation that are generally dependent on altered patterns of gene expression.
- Peptide hormones such as insulin, epinephrine, etc., and steroid hormones such as estrogen, progesterone, testosterone (in higher animals, for example, mammals), and ecdysone (in insects) control gene expression.
- In higher animals, hormones are synthesized in various specialized secretory cells (i.e., endocrine cells) and are released into the bloodstream.
- The peptide hormones do not normally enter cells because of their relatively large size. Their effects appear to be mediated by receptor proteins located in target-cell membranes and by the intracellular levels of cyclic AMP (cAMP) (called secondary messenger).
- The cAMP activates a protein kinase (for example., A-kinase) which phosphorylates (activates) many specific enzymes.
- The steroid hormones, on the other hand, are small molecules that readily enter cells through the plasma membrane.
- Once inside the appropriate target cells, the steroid hormones become tightly bound to specific receptor proteins which are present only in the cytoplasm of target cells.
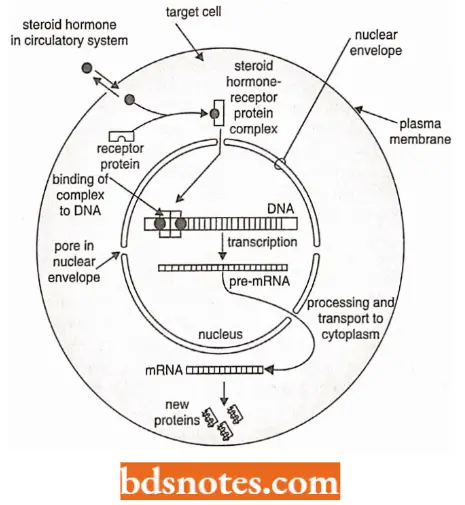
“Early warning signs of gaps in understanding gene regulation basics: Common questions”
- The hormone-receptor protein complexes activate the transcription of specific genes or sets of genes according to the following two methods (=hypotheses):
- The hormone receptor protein complexes interact with specific nonhistone chromosomal proteins and this interaction stimulates the transcription of the correct genes(J. Stein, G Stein, and L. Kleinsmith, 1975).
- The hormone receptor protein complexes activate the transcription of target genes by binding to specific DNA sequences present in the cis-acting regulatory regions (the enhancers and promoter regions) of the genes(R.M. Evans, 1988).
- In both of these cases, these hormone-receptor protein complexes would function as positive regulators (or “activators”) of transcription much like the CAP-cAMP complexes in prokaryotes.
- During the development of dipteran flies(Drosophila melanogaster and Chironomus tents), the steroid hormone ecdysone is released and triggers molting.
- If larvae of these insects are treated with ecdysone at stages of development before or between moultings, patterns of chromosome puffing occur that are identical to those occurring during natural moultings. Everyone tends to affect gene expression at the level of transcription.
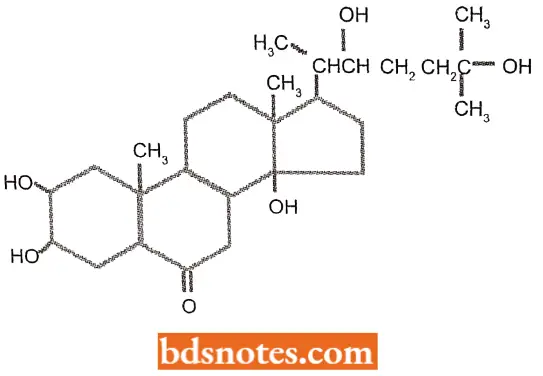
“Asymptomatic vs symptomatic effects of ignoring eukaryotic gene regulation principles: Q&A”
Regulation Of Gene Action Multiple Choice Questions And Answers
Question 1. Methylation of DNA most commonly occurs in the sequence of
- Cp G
- Cp A
- Cp T
- CpC
Answer: 1. Cp G
Question 2. The “Gene-battery model” of gene regulation in eukaryotes is proposed by
- Jacob and Monod
- Britten and Davidson
- Beadle and Tatum
- Komberg and Ochoa
Answer: 2. Britten and Davidson
“Differential applications of promoter regions vs enhancer elements: Questions answered”
Question 3. Fisher and Krebs got the Nobel Prize in 1992 for their discovery of
- Organ and cell transplantation
- Reversible protein phosphorylation as a biological regulatory mechanism
- Single ion channels
- All of the above
Answer: 2. Reversible protein phosphorylation as a biological regulatory mechanism
Leave a Reply