Transfer Rna (tRNA)
The RNA which possesses the capacity to combine specifically with only one amino acid in a reaction mediated by a set of amino acid-specific enzymes called aminoacyl-tRNA synthetases; transfers that amino acid from the amino acid pool to the site of protein synthesis and recognizes the codons of the mRNA is known as the soluble RNA (sRNA) or transfer RNA (tRNA).
- Thus, the tRNA molecule has to perform several highly complex functions during protein synthesis-it interacts with a specific synthetase enzyme, and possesses a site for binding an amino acid.
- Possesses a second site for interacting with a ribosome, and contains an anticodon that must be exposed to the codons of mRNA.
- Structure of tRNA. Robert Holley (1965) and his colleagues reported the complete nucleotide sequence of alanine tRNA of yeast (Holley received the Nobel Prize in 1968 for his work along with Khorana and Nirenberg).
- Nucleotide sequences are now known for more than 100 different “species” of tRNA.
“Understanding tRNA through FAQs: Q&A explained”
Transfer RNA has several unique characteristics:
- It is a relatively small molecule of 75 to 90 ribonucleotides and is, thus, smaller than either mRNA or any of the rRNAs, and has a sedimentation coefficient of 4S.
- The ratios of A: U and G: C are near unity which suggests the formation of DNA-like double helical segments (secondary structure).
- In these double-helical segments, G: C base pairs are more common than A: U as suggested by the ratio AU: GC = 0.7
- All tRNA molecules have a tertiary structure, the details for which are now known, and Mg ion concentration is important for its stabilization.
- Several Several “unusual” nucleotides are found in tRNA (for example., pseudouridineψ or psi), inosine Dihydropyridine (DHU), etc.). Many “unusual” nucleotides are methylated derivatives of common ones (for example., 1-methylguanylic acid, 1-methyl adenylic acid, ribothymidylic acid, and 5- methylcytosine).
The significance of these unusual bases of tRNA was understood well by molecular biologists during the construction of a two-dimensional model from the primary sequences of nucleotides of known tRNA.
- Thus, it was realized that most bases of tRNA pair according to Watson-Crick’s pairing rule, but unusual bases fail to do so because they carry substitutions or alterations in those positions that usually participate in hydrogen bonding.
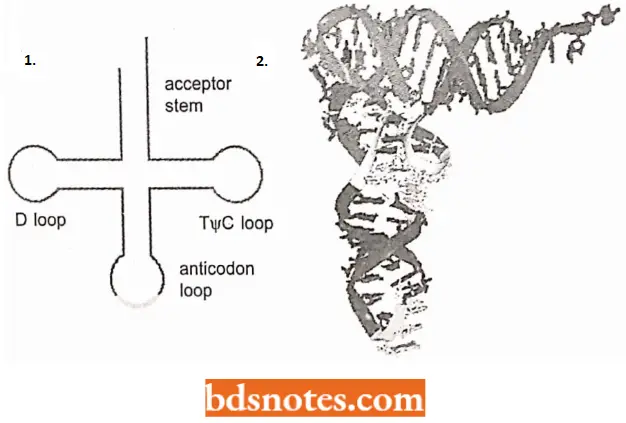
- Consequently, the presence of these bases forces the model builder to construct several non-base-paired loops in the tRNA molecule. By working on these lines, R. Holley (1965) first of proposed a clover leaf model for yeast tRNAala.
- The clover leaf model of tRNA accommodated several of the known functions of tRNA, so, it gained general acceptance.
“Importance of studying tRNA for biology students: Questions explained”
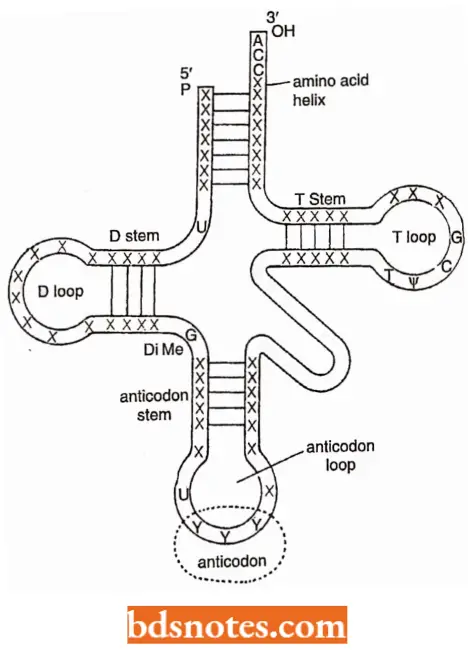
A Typical Clover-leaf Model tRNA Depicts The Following Structural Peculiarities:
- All tRNA molecules have guanine residue G at the 5′ terminal end and unpaired (single-stranded) C-C-A sequence at the 3′ end.
- This is called the amino acid attachment site because the amino acid becomes covalently attached to adenylic acid or A of the CCA sequence during polypeptide synthesis.
- The amino acid stem or helix consists of seven paired bases.
- The T-stein is composed of five paired bases-the last (i.e., nearest the T-loop or T\|/C loop is C-G.
- T-loop contains seven unpaired bases and is involved in binding rRNA molecules to the ribosomes.
- The anticodon stem includes five paired bases.
- The anticodon loop consists of seven unpaired bases, the third, the fourth, and the fifth of which (from the 3’) end of the molecule) constitute the anticodon.
- The anticodon permits temporary complementary pairing with three bases (triplet codon) on mRNA.
- The base on the 3′ side of the anticodon is a purine. Immediately adjacent to the 5′ side of the anticodon, uracil and another pyrimidine occur.
- A purine, often dimethylguanylic acid, is located in the “corner” between the anticodon stem and the D stem.
- The D-stem is composed of three or four base pairs (depending on the “species” of tRNA).
- The DHU-loop or D-loop is also variable in size containing 8 to 12 unpaired bases. The D-loop helps in the binding of amino-acyl synthetase. The extra ar m is variable in nucleotide composition and is lacking entirely in some tRNA.
“Common challenges in understanding tRNA effectively: FAQs provided”
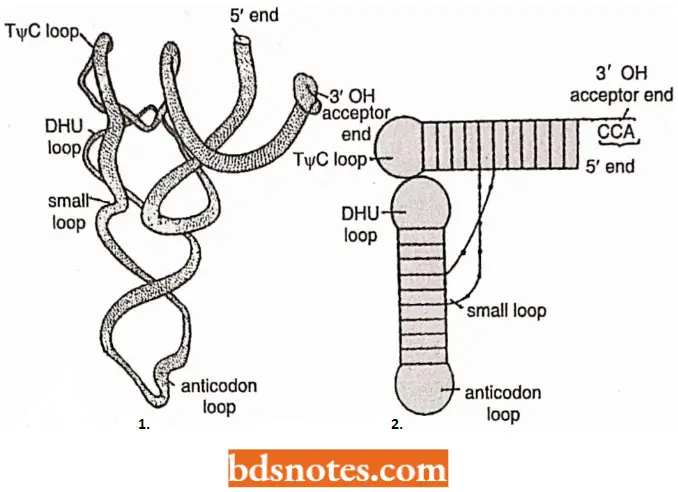
- Three-dimensional structure of tRNA. To understand the structure-function relationship of tRNA, its three-dimensional structure (TDS) was worked out with the help of an X-ray crystallography study.
- A. King, the Nobel laureate of 1982, has contributed much to the TDS of tRNAs.
- S.H. Kim (1973) proposed a most acceptable TDS model of tRNA of yeast cells). According to Kiin, the TDS of tRNA takes the shape of the letter L with a thickness of 10A°.
- Each arm of the L doubled over by bonds holding complementary bases together. Such an L-shape can also easily be derived from two two-dimensional clover-leaf models.
“Why is early learning of tRNA critical for molecular biology? Answered”
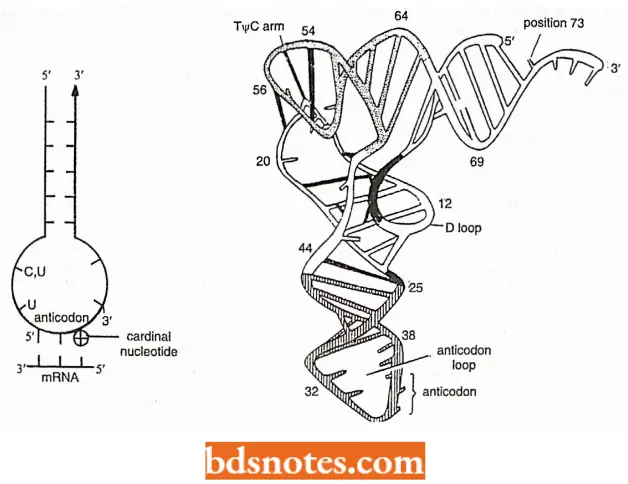
- Extended anticodon hypothesis. Recently it has been reported that the performance of anticodon, when isolated from tRNA, is weak and inaccurate.
- However, the performance of this anticodon triplet is enhanced if a matching sequence is present on the anticodon loop and the stem on either side of the anticodon triplet.
- As a result, an extended anticodon hypothesis has been proposed which suggests that the structures of the anticodon loop and that of the proximal anticodon stem are related to the sequence of anticodon (Michael Yarns, 1982).
Thus, the anticodon is extended into the nearby sequence and consists of all 12 nucleotides arranged in the following way:
- Two nucleotides (z.e., CU,ψU or UU) at the 5′ side of the anticodon loop;
- Three nucleotides of anticodon (3′ nucleotide of anticodon being very important and termed cardinal nucleotide);
- Two nucleotides at the 3′ side of the anticodon loop; and (zv) five pairs of nucleotides in the anticodon stem which can be conveniently written by giving only the bases on the 3′ side of the stem.
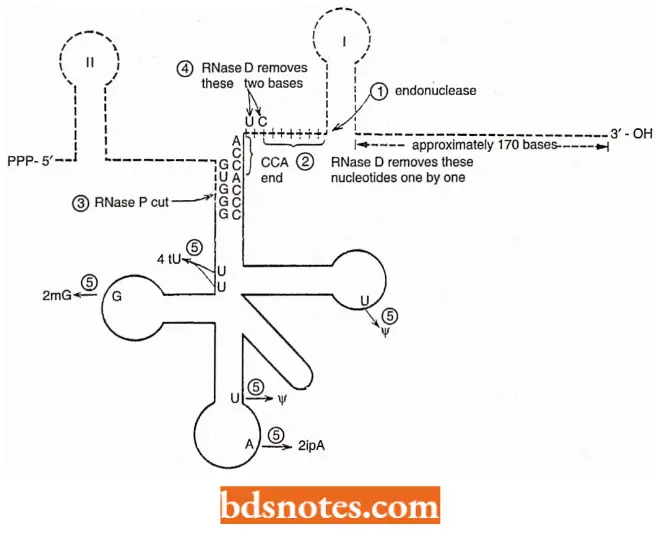
Processing of tRNA. Transfer RNA is transcribed from several particular sites on template DNA and comprises about 15 percent of the RNA present at any one time in an E. coli cell.
- Like the synthesis of rRNA and mRNA, the synthesis of tRNA molecules is initiated at a promoter site and completed at a terminator sequence. Transfer RNAs are also processed from larger precursors.
- In prokaryotes, tRNA processing includes the removal of nucleotides from the precursor or primary transcript and modification of some internal nucleotides.
- For example, in E.coli, the precursor for the amino acid tyrosine (pre tRNAtyr) consists of 128 bases; in processing to tRNAtyr 41 nucleotides are removed from the 5′ end and two are removed from the 3’end.
“Asymptomatic vs symptomatic effects of ignoring tRNA principles: Q&A”
In addition to processing, methylation, and inclusion of other “unusual” intercalary bases takes place after transcription, as does the addition of the 3′ terminal -C-C-A.
- The final functional tRNA molecule has 85 bases.
- A three-dimensional shape is achieved through hydrogen bonding.
- The processing of eukaryotic tRNAs resembles that of prokaryotes. However, in them, it is far more complex.
“Steps to explain tRNA structure: Anticodon vs amino acid attachment site: Q&A guide”
Splicing Of tRNA Precursors: The tRNA precursor splicing reaction has been worked out in detail in the yeast Saccharomyces cerevisiae.
- Both in vitro splicing systems and temperature-sensitive splicing mutants have been used in dissecting the tRNA splicing mechanism in S. cerevisiae.
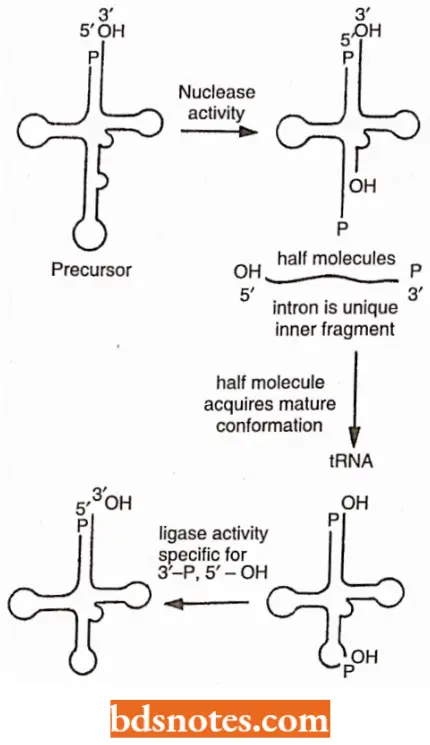
The excision of introns from yeast tRNA precursors occurs in two stages.
- In the first step, a nuclear membrane-bound splicing endonuclease enzyme makes two cuts precisely at the ends of the intron.
- Then, in a fairly complex set of reactions, a splicing ligase enzyme joins the two halves of the tRNA to produce the mature form of the tRNA molecule.
The specificity of these reactions exists in the conserved three-dimensional structural features of the tRNA precursors, not in the nucleotide sequences per sc (i.e., by itself). Cleavage of the tRNA precursor yields 5′ – OH termini and 2′ – 3′ cyclic phosphate groups at the 3′ termini.
The Stage 2 Ligation Process Involves Four Separate Reactions: The first reaction is the addition of a phosphate group to the 5′-OH terminus; this reaction requires kinase (enzyme) and a phosphate donor (ATP).
- The second reaction, the 5′ phosphate group is activated by the transfer of an AMP group to the terminus from an AMP-ligase intermediate (the AMP originally having been derived from ATP also).
- The 2′- 3’cyclic phosphate is opened by a cyclic phosphodiesterase activity that produces a 2’- phosphate and a free 3′ hydroxyl.
- The final ligation reaction occurs via a nucleophilic attack of the free 3′- OH on the interior
5’phosphate with the release of AMP. - All four of these reactions are catalyzed by the splicing ligase.
- Finally, the 2′- phosphate group (remaining from the 2′- 3’cyclic phosphate produced by the original cleavage reaction) is removed by a phosphatase activity to yield the mature tRNA molecule.
“Early warning signs of gaps in understanding tRNA basics: Common questions”
The overall two-stage mode of tRNA intron excision appears to occur in other organisms as well, for example., plants, mammals, etc. (Gardner et al., 2002).
- Earlier we considered a simple definition of a gene as a length of DNA that codes for one protein.
- But we have just encountered a variance: genes code for both tRNAs and rRNAs, yet neither is eventually translated into a protein. Their transcription function as final products without ever being translated.
- Thus, tRNA and rRNA are the major exceptions to the general rule that a gene codes for a protein.
- Genes for tRNA. There are, probably, at least 30 to 40 different tRNA genes and tRNA molecules in E. coli.
Higher organisms are found to contain 60 tRNA molecules and 60 tRNA genes. Since the cell uses only 20 amino acids in protein synthesis (and probably only 20 synthetase enzymes), it follows that several tRNAs will often have an affinity for the same amino acid.
- For example, E. coli cells contain five species of tRNA for leucine amino acid.
- All the tRNA genes constitute far less than 1% of the total genome in both E.coli and eukaryotic cells, yet some 10 to 15% of each cell’s RNA may be in the form of tRNA.
This discrepancy between the number of tRNA genes and gene transcripts occurs because of the following facts-
- The tRNA molecules are relatively stable compared with many kinds of RNA.
- The tRNA molecules are transcribed continuously and more quickly by tRNA genes than other RNAs because they are needed in plentiful amounts.
“Role of tRNA in translation: Questions answered”
Leave a Reply