RNA Editing
In the late 1980s, still, another quite unexpected form of post-transcriptional RNA processing was discovered in several organisms.
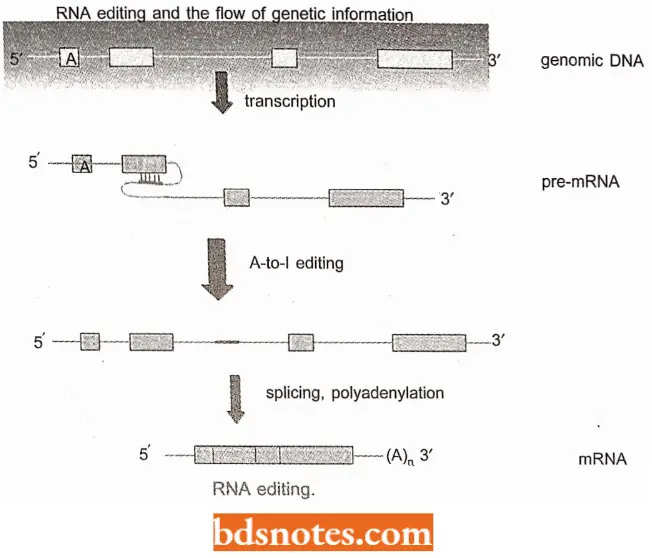
- In this form, referred to as RNA editing, the nucleotide sequence of a pre-mRNA is changed before translation.
- As a result, the ribonucleotide sequence of the mature RNA differs from the sequence encoded in the exons of the DNA from which the RNA was transcribed.
RNA Editing Is Of The Following Two Types:
- Substitution RNA editing;
- Insertion or deletion RNA editing.
“Asymptomatic vs symptomatic effects of ignoring RNA editing principles: Q&A”
“Understanding RNA editing through FAQs: Q&A explained”
Substitution RNA Editing: In this process, the identities of individual nucleotide bases are altered.
- Substitution editing is used in some nuclear-derived eukaryotic RNAs and is also very prevalent in mitochondrial and chloroplast RNAs transcribed in plants.
Insertion Or Deletion RNA Editing: In this process, nucleotides are added to or subtracted from the total number of bases.
Examples Of RNA Editing
- The slime mold, Physarum polycephalum uses both substitution and insertion/deletion editing for its mitochondrial mRNAs.
- RNA editing was particularly evident in mitochondrial proteins of a group of protozoan parasites, the trypanosomes (some of which cause African sleeping sickness); in one case, more than 50% of the nucleotides in the mRNA were added uridines (Us).
“Early warning signs of gaps in understanding RNA editing basics: Common questions”
Uridines were also deleted from the original sequence. These parasites had another curious trait- the existence of minicircles and maxicircles of DNA in specialized mitochondria, called kinetoplasts.
- In the average kinetoplasts, there are about 50 maxicircles and about 5000 minicircles, concentrated like chain links.
- The maxicircles contain genes for mitochondrial function.
- Larry Simpson and his colleagues showed in 1990, that both maxicircles and minicircles are templates for guide RNA (gRNA), RNA that guides the process of messenger editing.
- The guide RNA forms a complement with the mRNA to be edited; however, the gRNA has a sequence complementary to that of the final mRNA, the one with bases added.
- Since the bases have not yet been added, a bulge occurs in the gRNA where the complement to be added is.
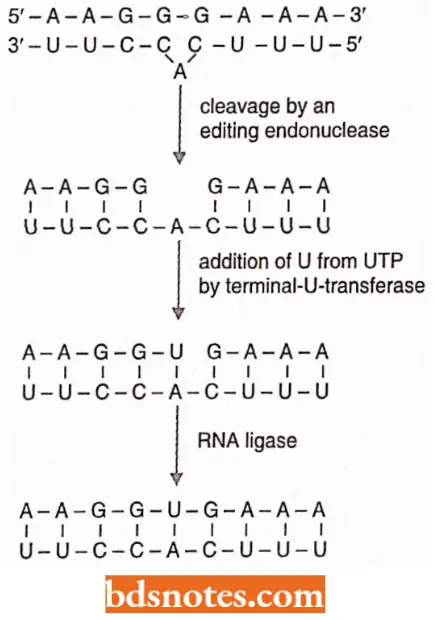
- The mRNA is then cleaved opposite the bulge by an enzyme-editing endonuclease.
A uridylate (U) is brought into the mRNA as a complement to the adenine with the enzyme terminal-U-transferase. An RNA ligase then closes the nick in the mRNA, which now has a uridylate added.
“Importance of studying RNA editing for biology students: Questions explained”

An exciting outcome of this research, aside from learning about a novel mechanism of messenger RNA processing, is the possibility of clinical rewards.
- Anytime there is a specialized pathway in a parasite not found in its host, it is possible to use that pathway to attack the parasite.
- Thus, this research might lead to new approaches to combating these trypanosome parasites.
- The most well-studied examples of institutional editing occur in mammalian nuclear-encoded mRNA transcripts.
- For example, apolipoprotein-B (apo-B) exists in both a long and a short form though a single gene apoB encodes both proteins (Hodges and Scott 1992).
- In human intestinal cells, apo B mRNA is edited by a single C (called nucleotide 6666) to U change, which converts a CAA glutamine codon into a UA stop codon and terminates the polypeptide at approximately half its genomically encoded length.
- The editing is performed by a complex of proteins that bind to a “mooring sequence” (moor means tie up) on the mRNA transcript just downstream of the editing site.
“Role of RNA editing enzymes like ADARs and APOBECs: Questions answered”
“Common challenges in understanding RNA editing effectively: FAQs provided”
- The synthesis of subunits constituting the glutamate receiptor channels (GIu R) in mammalian brain tissue is also affected by RNA editing. In this case, adenosine to inosine (I) editing occurs in pre-mRNAs before their translation, during which I is read as guanosine (G).
- The enzymes produced by a family of three ADAR (Adenosine Deaminase Acting on RNA) genes are believed to be responsible for the editing of various sites within the glutamate channel subunits.
- The double-stranded RNAs required for editing by the ADAR enzymes are provided by the intron-exon pairing of the GluR mRNA transcripts.
- The editing changes the physiological parameters (i.e., solute permeability and desensitization response time) of the receptors containing the subunits.
The importance of RNA editing resulting from the action of ADARs is most apparent when one examines situations in which these enzymes have lost their functional capacity as a result of mutation, in several investigations, the loss of function was shown to have a lethal impact on mice.
“Steps to explain RNA editing mechanisms: Adenosine-to-inosine (A-to-I) vs cytidine-to-uridine (C-to-U): Q&A guide”
- In one study, embryos heterozygous for a defective ADAR1 gene die during embryonic development as a result of a defective hemopoietic system (Wang et al 2000).
- In another study, mice with two defective copies ofADAR2 progress through development normally, but are prone to epileptic seizures and die while still in the weaning stage. Their tissues contain the unedited version of one of the Glu R products.
- The defect to death is believed to be in the brain. Heterozygotes for the mutation are normal.
Leave a Reply