Nucleosides And Nucleotides
The nucleic acids DNA (deoxyribonucleic acid) and RNA (ribonucleic acid) are the molecules that play a fundamental role in storing genetic information, and the subsequent manipulation of this information. They are polymers whose building blocks are nucleotides, which are themselves combinations of three parts: a heterocyclic base, a sugar, and phosphate.
The most significant difference in the nucleotides comprising DNA and RNA is the sugar unit, which is deoxyribose in DNA and ribose in RNA. The term nucleoside is used to represent a nucleotide lacking the phosphate group, i.e. the base–sugar combination.
The general structure of nucleotides and nucleosides is shown below:
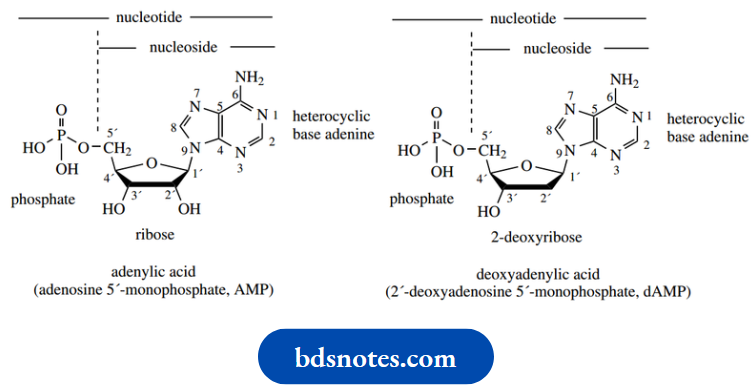
Before we analyze nucleotide structure in detail, it is perhaps best that we consider the nature of the various parts. In nucleic acid structures, there are five different bases and two different sugars.
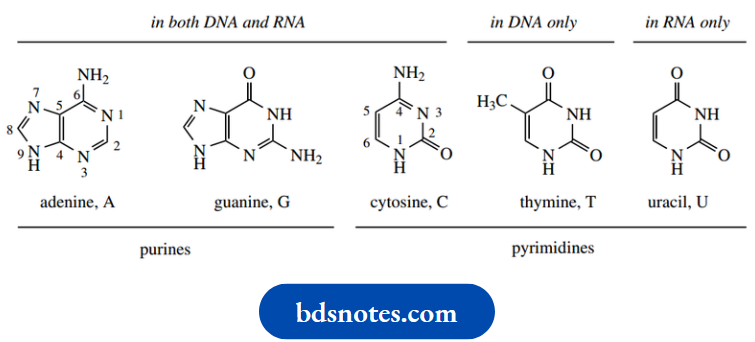
The bases are monocyclic pyrimidines (see or bicyclic purines and all are aromatic. The two purine bases are adenine (A) and guanine (G), and the three pyrimidines are cytosine (C), thymine (T) and uracil (U). Uracil is found only in RNA, and thymine is found only in DNA. The other three bases are common to both DNA and RNA. The heterocyclic bases are capable of existing in more than one tautomeric form. The forms shown here are found to predominate in nucleic acids. Thus, the oxygen substituents are in keto form, and the nitrogen substituents exist as amino groups.
Sugars:
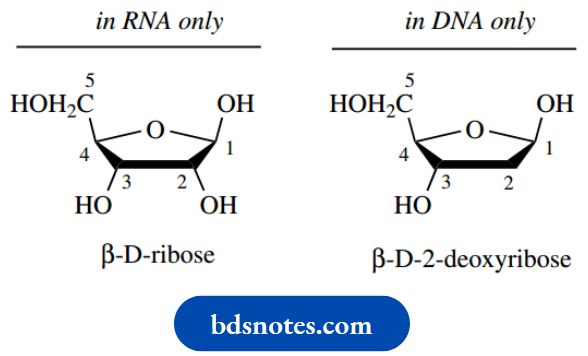
The two sugars are pentoses, D-ribose in RNA and 2-deoxy-D-ribose in DNA. In all cases, the sugar is present in a five-membered acetal ring form, i.e. a furanoside. The base is combined with the sugar through an N-glycoside linkage at C-1, and this linkage is always β.
Purine bases are linked through N-9, and pyrimidines through N-1. When numbering nucleosides and nucleotides, we use primed numbers for the sugar, since non-primed numbers are already employed in the base part. There are thus four different nucleosides for each type of nucleic acid, as shown.
Nucleosides in RNA:
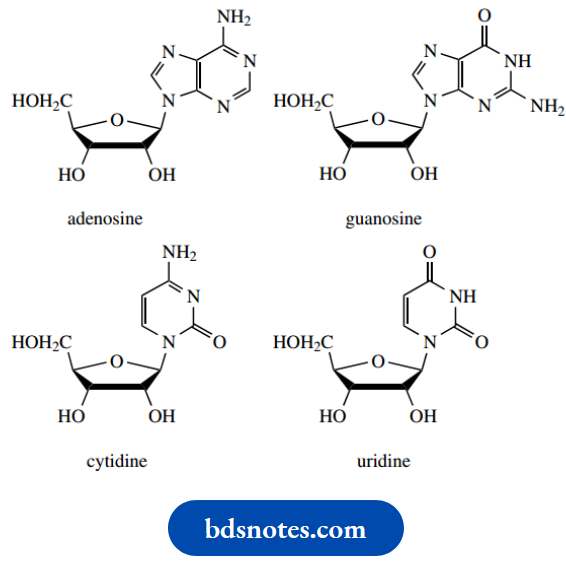
Nucleosides in DNA:
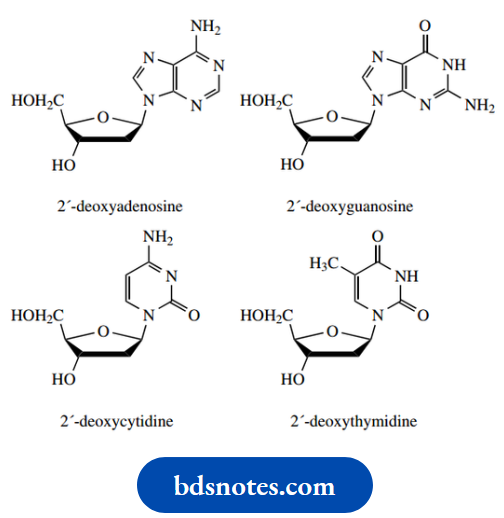
The phosphate group of nucleotides is attached via a phosphate ester linkage, and may be attached to either C-5′ or C-3′. As we shall see, nucleosides in nucleic acids are joined together through a phosphate linkage between the 3′-hydroxyl of one sugar and the 5′-hydroxyl of another. As a result, hydrolysis of nucleic acid could give us nucleotides containing either 5′- or 3′-phosphate groups. It is usual, however, to consider nucleic acids as composed of nucleotides containing a 5′-phosphate group.
The accepted nomenclature for the various components in RNA and DNA
Nomenclature of bases, nucleosides, and nucleotides:
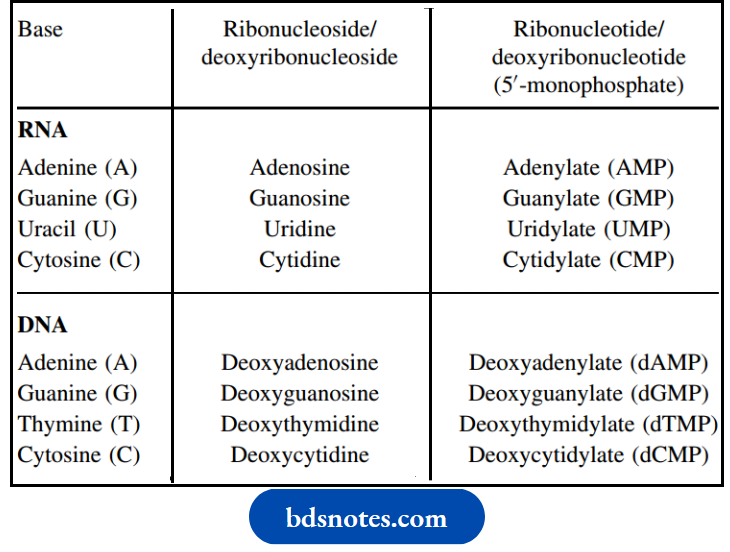
Nucleic Aids
1. DNA
The nucleic acids comprise a long unbranched chain of nucleotide monomeric units. The nucleotides are linked together via the phosphate group, which joins the sugar units through ester linkages, usually referred to as phosphodiester bonds.
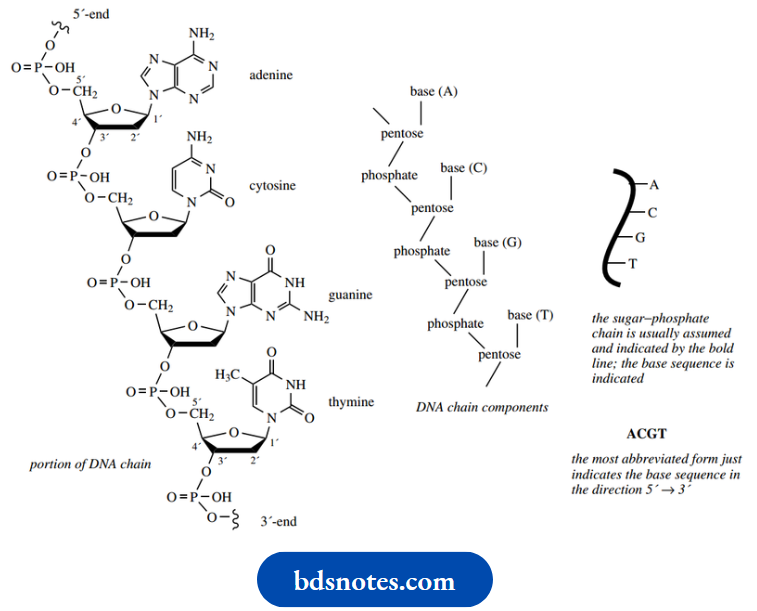
- The phosphodiester bond links the 5′ position of one sugar with the 3′ position of the next. A short portion of a DNA molecule
- The nucleic acid chain is thus composed of alternating units of sugar and phosphate, with the bases appearing as side chains from the sugar components. The nucleotide chain has ends, referred to as the 5′- and 3′-ends, according to the sugar hydroxyl that is available for further bonding.
- Though we shall not be considering this aspect further, in some organisms, especially bacteria and some viruses, the two ends of the DNA chain are joined together so that we encounter a circular form of DNA.
- Nucleic acid structures generally need to be written in a much-abbreviated form. The sugar-phosphate backbone is taken for granted; it can be indicated by a line, with the attached bases defined.
- Even this is tedious. It is thus reduced further to the sequence of attached bases. The base sequence of the nucleic acid is the standard way of defining its structure; strictly, the structure is a sequence of nucleotides.
- By convention, the base sequence is written from the 5′-end to the 3′-end, so that the short strand of DNA would be given as–ACGT–
Perhaps the most far-reaching feature of nucleic acids is the ability of the bases to hydrogen bond to other bases. This property is fundamental to the double helix arrangement of the DNA molecule, and the translation and transcription via RNA of the genetic information present in the DNA molecule.
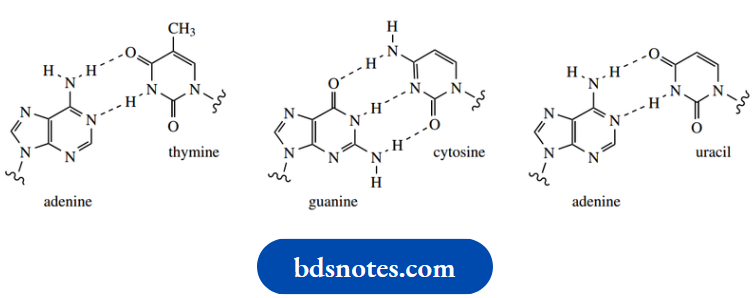
The polymeric strand of DNA coils into a helix, and it is bonded to a second helical strand by hydrogen bonds between appropriate base pairs. In DNA, the base pairs are adenine–thymine and guanine-cytosine. It should be appreciated that each of these bases is planar and that the hydrogen-bonded base pair is also planar. The hydrogen-bonded N–H–N and N–H–O interatomic distances are in the range 2.8–3.0 A. By comparison, N–H and O–H˚ bonds are typically about 1.0 Å.
Thus, each purine is specifically linked to a pyrimidine by either two or three hydrogen bonds. The result of these interactions is that each nucleotide recognizes and bonds with its complementary partner.
This specific base pairing means that the two strands in the DNA double helix are complementary. Wherever adenine appears in one strand, thymine appears opposite it in the other; wherever cytosine appears in one strand, guanine appears opposite it in the other. We shall see later the significance of base pairing between adenine and uracil. The latter base is found in RNA instead of thymine.
The DNA double helix has both chains twisting on a common axis. The bases are directed inwards to allow hydrogen bonding, and the sugar and phosphodiester parts of the main chain form the outside portion.
The planes of the base pairs are perpendicular to the helix axis so that the molecule looks like a spiral staircase with the base-pair combinations forming the treads.
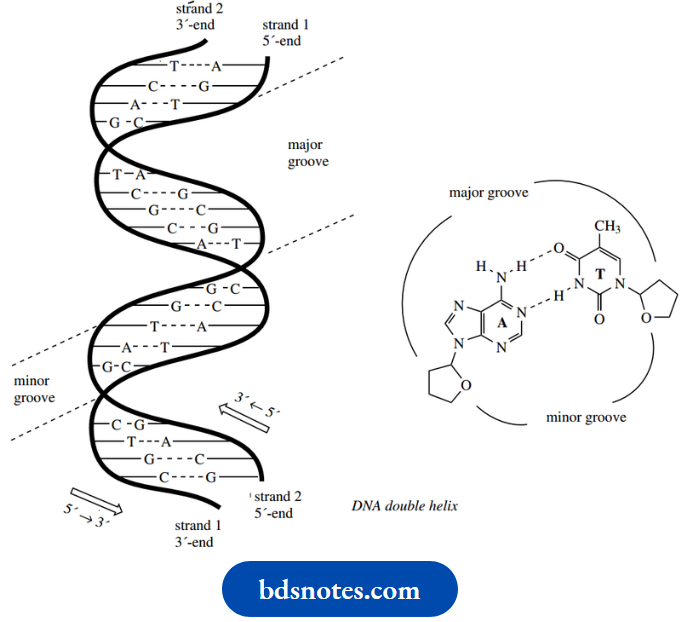
The helix makes A complete turn every 10 base pairs along the chain. The two strands are complementary:
If you know the sequence along one chain, you can write down the sequence along the other via the base pairing relationship. Note, however, that the chains are antiparallel, i.e. they run in opposite directions.
- This is indicated in the schematic diagram in
- One further point arises because the glycoside bonds between the sugars and bases of a particular base pair are not directly opposite each other.
- This is easily appreciated from the illustrations of hydrogen-bonded base pairings. The consequence of this is that the grooves along the outside of the double helix array are of unequal width, and are termed the major groove and the minor groove.
- These grooves contain many water molecules through interaction with amino and carbonyl groups of the bases, and are distinguishable to agents that bind to DNA,
Example: Some anticancer drugs.
2. Replication of DNA
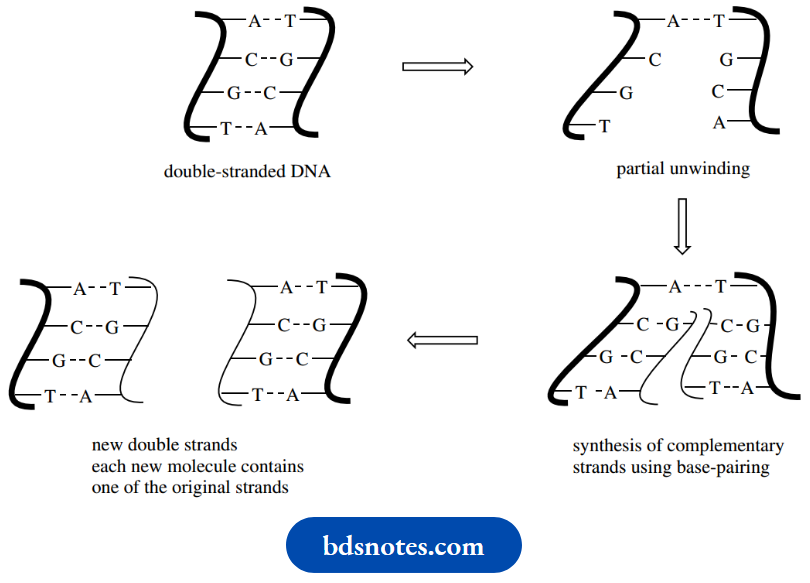
During cell division, the DNA molecule is replicated so that each daughter cell will carry its DNA molecule. During the process, the two strands of DNA unwind, and each strand then acts as the template for the synthesis of a new strand; in each case, the new strand is complementary to the original because of the base-pairing restrictions.
Each new double helix is comprised of one strand that was part of the original molecule and one strand that is newly synthesized. Not surprisingly, this is a very simplistic description of a quite complex process, catalyzed by enzymes known as DNA polymerases.
The precursors for the synthesis of the new chain are the nucleoside triphosphates, dATP, dGTP, dTTP, and dCTP. We have already met ATP when we considered anhydrides of phosphoric acid; these compounds are analogs of ATP, though the sugar is deoxyribose rather than ribose.
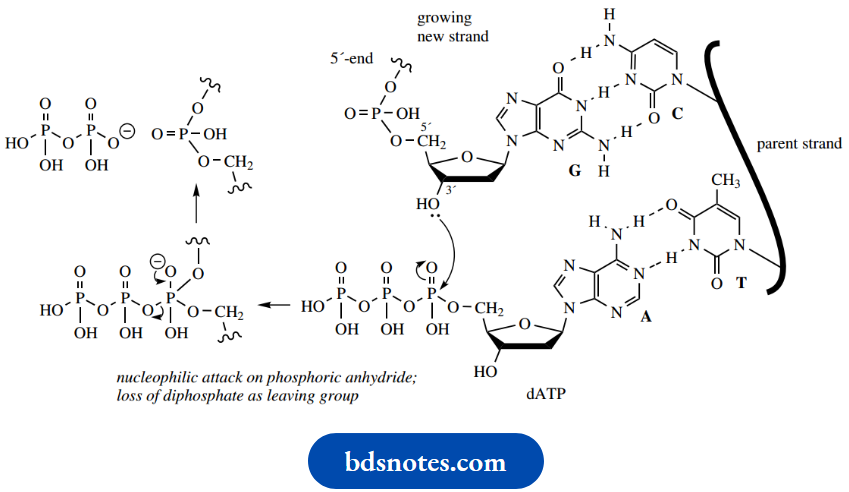
These triphosphate anhydrides are susceptible to nucleophilic attack by hydroxyl groups. Chain extension is simply an esterification reaction utilizing the 3′-hydroxyl of the sugar in the growing chain, with diphosphate as a good leaving group.
- The correct nucleoside triphosphate is selected because of the hydrogen bonding properties of base pairs. This also provides the correct alignment so that the reaction can occur.
- In the illustration, the next base in the original DNA strand is thymine, which dictates that only an adenine nucleotide can hydrogen bond and form the complementary base pair.
- The esterification occurs, with the loss of diphosphate as the leaving group, and the new daughter strand is extended by one nucleotide.
- The process repeats as the enzyme moves on to the next position on the original DNA strand. Hydrolysis of diphosphate to two molecules of phosphate provides some of the driving force to facilitate the reaction.
3. RNA
RNA differs structurally from DNA in three important ways. First, as indicated above, the sugar in RNA is ribose, not 2-deoxyribose.
- Second, thymine is replaced by uracil, so that the four bases are adenine, uracil, guanine, and cytosine.
- The third difference is that RNA is usually single-stranded. Although an RNA molecule may be single-stranded, it does not exclude the possibility of partial double-stranded sequences being present. In such cases, the molecule doubles back on itself and coils up with a complementary base sequence elsewhere.
- Remember that complementary sequences now involve A–U rather than A–T hydrogen-bonding interactions.
- DNA stores the genetic information for a cell, but it is RNA that participates in the processes by which this information is used. RNA molecules are classified according to their function or cellular location.
Three major forms are found in prokaryotic cells:
- Messenger RNA (mRNA) carries genetic information from DNA to ribosomes, the organelles responsible for protein synthesis;
- Ribosomal RNA (rRNA) is an integral part of the ribosomes
- Transfer RNA (tRNA) carries the amino acid residues that are added to the growing peptide chain during protein synthesis.
4. The genetic code
It is the sequence of bases along one of the strands of the DNA molecule, the coding strand, that provides the information for the synthesis of proteins, especially enzymes, in an organism.
- A complementary sequence exists along the second strand, and this is termed the template strand. A gene is a segment of DNA that contains the information necessary for the synthesis of one protein.
- Each amino acid in a protein is specified by a sequence of three nucleotides, termed a codon.
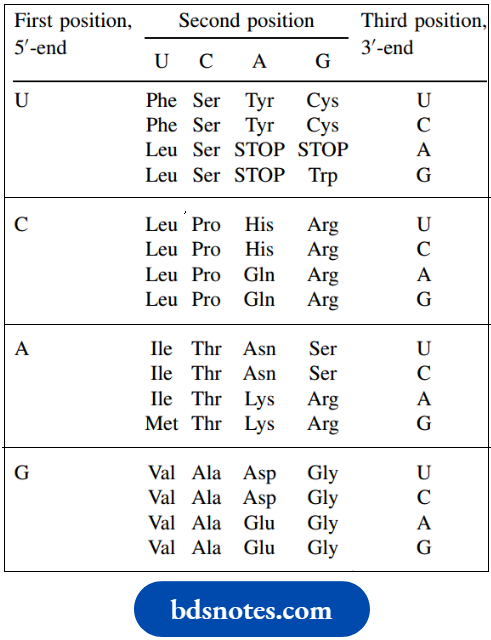
- A codon is usually designated in terms of the base sequence, however, just as we saw with nucleic acid sequences above. With four different bases, there are 43 = 64 different combinations of three bases (codons) available, more than enough for the 20 different amino acids found in proteins.
- Most amino acids can be specified by two or more different codons, and three particular codons are known to carry the signal for stop, i.e. chain termination.
- The signal for start is the same as for methionine (unusual in having only one codon rather than several) and means that all proteins should begin with a methionine residue.
- Since this is not the case, the inference is that many proteins are subsequently modified by cleaving off a fragment that contains this starter amino acid residue.
The codon combinations A codon can be the DNA sequence in the coding strand or the related sequence found in mRNA.The table shows the mRNA sequences since we shall be using these during consideration of protein synthesis. The DNA sequences merely have thymine (T) in place of uracil (U), as appropriate. The sequence is always listed from the 5′-end to the 3′-end.
The genetic code: mRNA sequences:
Transcription of DNA to mRNA:
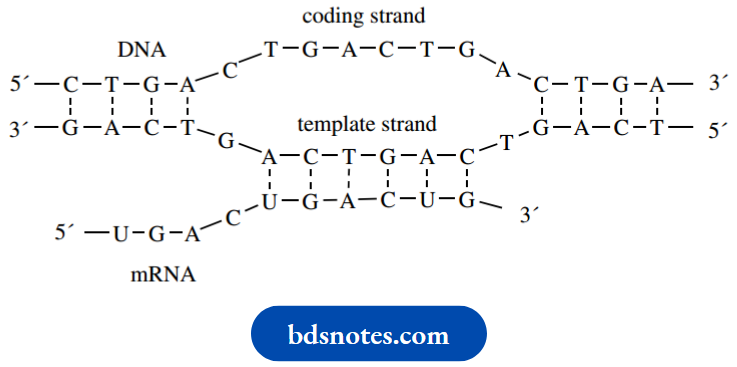
5. Messenger RNA synthesis transcription
Although the amino acid sequence of a protein is defined by the sequence of codons in DNA, it is RNA that participates in the interpretation of this sequence and the subsequent joining together of amino acids. The process starts with the synthesis of mRNA, in a process called transcription.
- Part of the DNA double helix, corresponding to the gene in question, is unwound. Rather like in the replication of DNA, the base sequence is used to synthesize a new nucleic acid strand.
- However, this time only one strand is interpreted, the template strand, and ribonucleotides (ATP, GTP, CTP, and UTP) are used in the new chain assembly instead of deoxyribonucleotides.
- The sequence of ribonucleotides incorporated is dictated by the sequence of nucleotides in DNA and depends on hydrogen bonding between pairs of bases.
- In RNA synthesis, uracil nucleotides are employed rather than thymine nucleotides. The result is a synthesis of a single strand of RNA with a sequence analogous to the coding strand of DNA, except that U replaces
- T. Coupling of the ribonucleotide units is catalyzed by the enzyme RNA polymerase and is mechanistically the same as with DNA replication above, i.e. esterification of a hydroxyl via a phosphoric anhydride.
6. Transfer RNA and translation
Although messenger RNA is synthesized in the cell nucleus, it then moves to the cytoplasm and to the ribosomes, where protein biosynthesis occurs.
- These particles are composed of two subunits, termed 50S and 30S, and are combinations of rRNA and protein.
- The ribosomes are responsible for binding the two other types of RNA, mRNA (which contains the genetic code) and tRNA (which carries the individual amino acids).
- tRNA molecules are very small compared with the other forms of RNA, being less than 100 nucleotides.
- The size of mRNA reflects the number of amino acid residues in the protein being synthesized but could be a thousand or more nucleotides. rRNA is the most abundant of the three types of RNA, and in size covers a range from about 75 to 3700 nucleotides.
- A tRNA molecule is specific for a particular amino acid, though there may be several different forms for each amino acid. Although relatively small, the polynucleotide chain may show several loops or arms because of base pairing along the chain.
- One arm always ends in the sequence cytosine–cytosine–adenosine. The 3’hydroxyl of this terminal adenosine unit is used to attach the amino acid via an ester linkage.
However, it is now a section of the nucleotide sequence that identifies the tRNA–amino acid combination, and not the amino acid itself.
- A loop in the RNA molecule contains a specific sequence of bases, termed an anticodon, and this sequence allows the tRNA to bind to a complementary sequence of bases, a codon, on mRNA.
- The synthesis of a protein from the message carried in mRNA is called translation, and a simplified representation of the process as characterized in the bacterium.
- Escherichia coli is shown below. Initially, the amino acid is activated by an ATP-dependent process, producing an aminoacyl-AMP.
- A hydroxyl group in ribose, part of a terminal adenosine group of tRNA, then reacts with this mixed anhydride. In this way, the amino acid is bound to tRNA via an ester linkage as an aminoacyl-tRNA.
The tRNA involved will be specific for the particular amino acid. A detailed mechanism for this process has been considered in
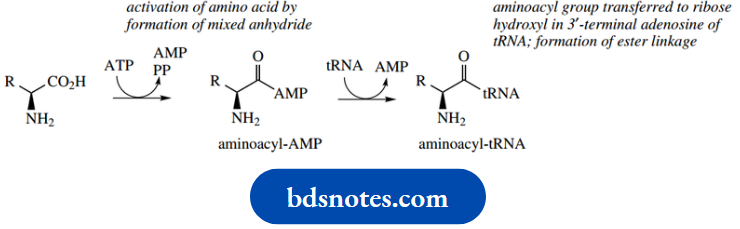
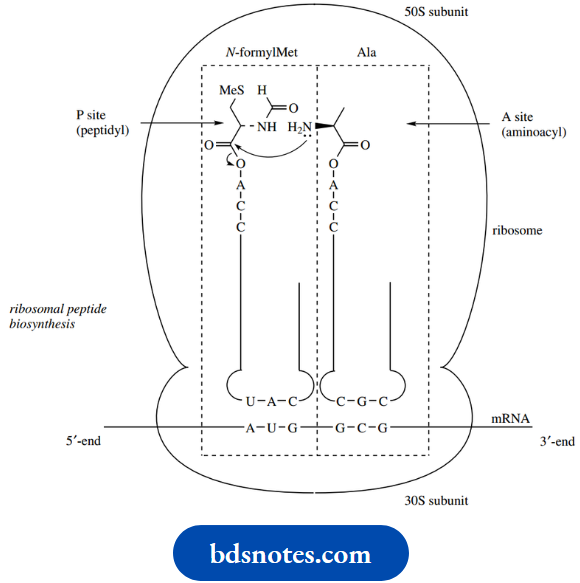
The mRNA is bound to the smaller 30S subunit of the bacterial ribosome. The mRNA is a transcription of one of the genes of DNA and carries the information as a series of three-base codons. The message is read (translated) in the 5’to 3’direction along the mRNA molecule.
- The aminoacyl-tRNA anticodon (UAC) allows binding via hydrogen bonding to the appropriate codon (AUG) on mRNA. In prokaryotes, the first amino acid encoded in the sequence is N-formylmethionine (fMet).
- Although the codon for initiation ( N-formylmethionine) is the same as that for methionine, the initiator tRNA used is different from that employed for the incorporation of methionine elsewhere in the peptide chain.
- The initiator aminoacyl-tRNA is thus bound and positioned at the P (for peptidyl) site on the ribosome. The next aminoacyl-tRNA a tRNA specific for alanine) is also bound via a codon (GCG)–anticodon (CGC) interaction and is positioned at an adjacent A (for aminoacyl) site on the ribosome.
This allows peptide bond formation to occur, with the amino group of the amino acid in the A site attacking the activated ester in the P site. The peptide chain is thus initiated and has become attached to the tRNA located in the A site.
The tRNA at the P site is no longer required and is released from the ribosome. Then the peptidyl-tRNA at the A site is translocated to the P site by the ribosome moving along the mRNA a codon at a time, exposing the A site for a new aminoacyl-tRNA appropriate for the particular codon, and a repeat of the elongation process occurs. The cycles of elongation and translocation continue until a termination codon is reached, and the peptide or protein is then hydrolyzed and released from the ribosome.
Note that the protein is synthesized from the N-terminus towards the C-terminus. Some special features of proteins are elaborated by secondary transformations that are not part of the translation process. The N-formylmethionine initiator may be hydrolyzed to methionine, or, as we have already indicated, the methionine unit may be removed altogether. Other post-translational changes to individual amino acids may be seen,
Example:
The hydroxylation of proline to hydroxyproline or the generation of disulfide bridges between cysteine residues
Antibiotics that interface with ribosomal peptide biosynthesis
Many of the antibiotics used clinically are active because of their ability to inhibit protein biosynthesis in bacteria. The individual steps of protein biosynthesis all seem susceptible to disruption by specific agents.
Some specific examples are listed below:
- Inhibitors of transcription
- Rifampicin(inhibits RNA polymerase)
- Inhibitors of aminoacyl-tRNA binding to ribosome
- Tetracyclines (bind to 30S subunit of the ribosome and prevent attachment of aminoacyl-tRNA)
- Inhibitors of translation
- Streptomycin (binds to 30S subunit of the ribosome, causes mRNA to be misread)
- Erythromycin (binds to 50S subunit of the ribosome, inhibits translocation)
- Chloramphenicol (binds to 50S subunit, inhibits peptidyltransferase activity)
Naturally, if such materials are going to be useful as antibiotic drugs, we require a selective action. We need to be able to inhibit protein biosynthesis in bacteria, whilst producing no untoward effects in man or animals. Although the mechanisms for protein biosynthesis are essentially the same in prokaryotes and eukaryotes, there are some subtle differences, e.g. the ribosome and how the process is initiated. Without such differences, the agent would be toxic to man as well as to bacteria.
Nucleosides as antiviral agents
Viruses are responsible for many human and animal diseases, with a variety of symptoms and levels of severity.
- Common viral illnesses include colds, influenza, cold sores (herpes), and childhood infections such as chickenpox, measles, and mumps.
- More serious conditions include meningitis, poliomyelitis, and human immunodeficiency virus (HIV), the latter potentially leading to acquired immune deficiency syndrome (AIDS).
- Viruses are simpler than bacteria and consist essentially of nucleic acid (either DNA or RNA) enclosed in a protein coat.
- Those causing chickenpox, smallpox, and herpes belong to the DNA virus group, whereas those responsible for influenza, measles, mumps, meningitis, poliomyelitis, and HIV are classified as RNA viruses. Viruses have no metabolic machinery of their own, and for their very existence are intracellular parasites of other organisms.
- To survive and reproduce, they have to tap into the metabolic processes of the host organism.
- For this reason, it is difficult to find drugs that are selective towards viruses without damaging the host.
- Most antiviral agents are only effective whilst the virus is replicating, and viral replication is very far advanced by the time the infection is detectable. There are relatively few effective antiviral drugs, and most of these are nucleoside derivatives.
Aciclovir
Aciclovir (acyclovir) was one of the first effective selective antiviral agents. It is a guanine derivative of value in treating herpes viruses, though it does not eradicate them, and is only useful if drug treatment is started at the onset of infection.
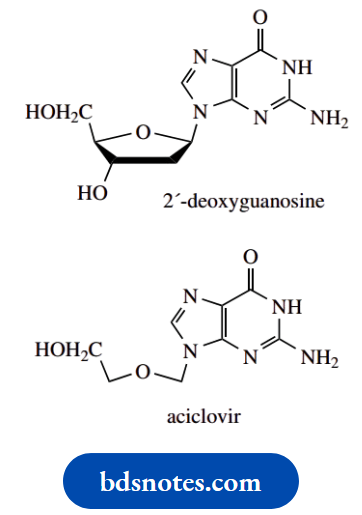
Aciclovir is a member of a group of nucleoside derivatives termed acyclonucleosides, in that there is an incomplete sugar ring.
- The structural relationship to 2′-deoxyguanosine should be very clear.
- Aciclovir is converted into its monophosphate by the viral enzyme thymidine kinase – some viruses also possess enzymes that facilitate their replication in the host cell.
- The viral enzyme turns out to be much more effective than that of the host cell, and conversion is, therefore, mainly in infected cells.
- The monophosphate is subsequently converted into triphosphate by the host cell enzymes.
- Aciclovir triphosphate inhibits viral DNA polymerase, much more so than it does the host enzyme, and so terminates DNA replication.
Zidovudine:
Zidovudine is 3′-azido-3′-deoxythymidine and is a derivative of deoxythymidine in which an azide group replaces the 3′-hydroxyl.
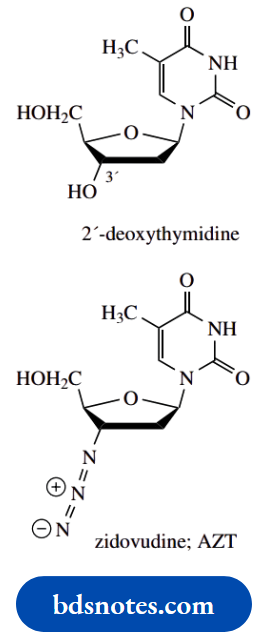
It is better nown as the anti-AIDS drug AZT.
- The AIDS virus is an RNA retrovirus. In retroviruses, an enzyme reverse transcriptase makes a DNA copy of viral RNA (contrast transcription: making an RNA copy of DNA).
- This DNA copy is then integrated into the host genome and gets transcribed into both new viral RNA and mRNA for translation into viral proteins. AZT is an inhibitor of reverse transcriptase.
- AZT is phosphorylated by cellular enzymes to the triphosphate, which competes with normal substrates for the formation of DNA by reverse transcriptase and blocks viral DNA synthesis.
- Mammalian DNA polymerase is relatively unaffected, but there can be some toxic effects. AZT is used in AIDS treatment along with other antiretroviral drugs.
Some Other Important Nucleosides And Nucleotides ATP SAm Coenzyme A, NAD, FAD
The terminology nucleotide or nucleoside immediately directs our thoughts toward nucleic acids.
- Remarkably, nucleosides and nucleotides play other roles in biochemical reactions that are no less important than their function as part of nucleic acids.
- We also encounter more structural diversity.
- It is rare that the chemical and biochemical reactivities of these derivatives relate specifically to the base plus sugar part of the structure, and usually reside elsewhere in the molecule.
Almost certainly, it is this base plus sugar part of the structure that provides a recognition feature for the necessary enzymes that utilize these compounds.
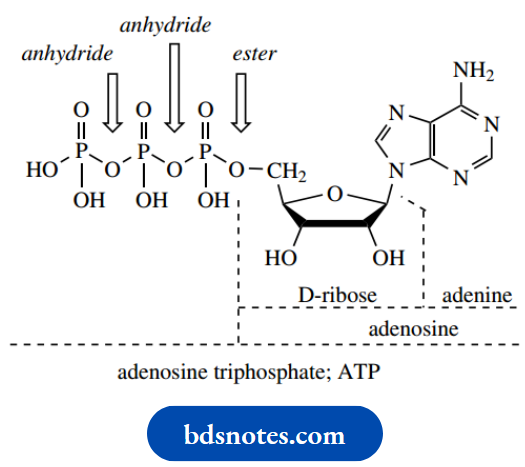
ATP:
ATP, adenosine triphosphate, provides the currency unit for energy in biochemical reactions (see and is simply a triphosphate variant of a standard RNA nucleotide. It is, of course, the biosynthetic precursor for adenine-based units in RNA. As we have already seen, the functions of ATP can be related to hydrolytic reactions in the triphosphate (anhydride) part of the molecule.
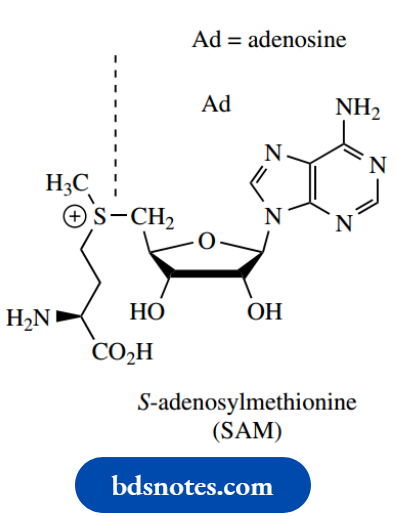
SAM:
SAM, S-adenosylmethionine, has been encountered as a biological methylating agent, carrying out its function via a simple SN2 reaction.
This material is a nucleoside derivative formed by nucleophilic attack of the thiol group of methionine onto ATP. It provides in its structure an excellent leaving group, the neutral Sadenosylhomocysteine.
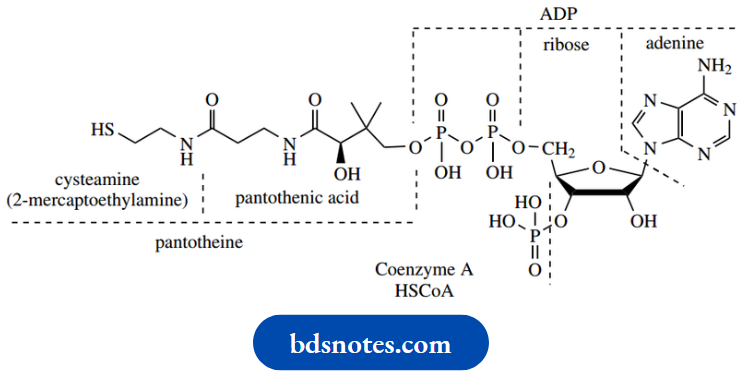
Coenzyme A
Coenzyme A is another adenine nucleotide derivative, with its primary functional group, a thiol, some distance away from the nucleotide end of the molecule. This thiol plays an important role in biochemistry via its ability to form thioesters with suitable acyl compounds.
We have seen how thioesters are considerably more reactive than oxygen esters, with particular attention being paid to their improved ability to form enolate anions, coupled with thiolates being excellent leaving groups.
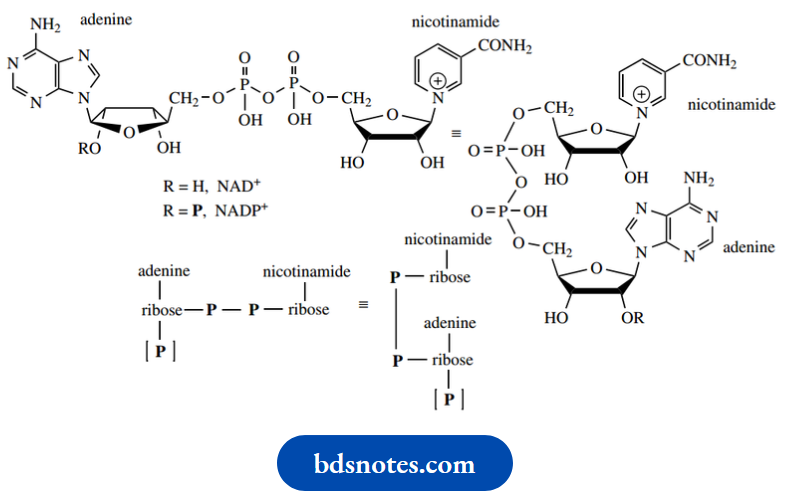
Nature’s oxidizing agents NAD+ and NADP+, and the corresponding reducing agents NADH and +, are all dinucleotide derivatives).
- Indeed, the full names betray this: NAD is nicotinamide adenine dinucleotide. From the structures of nucleic acids, one interprets a dinucleotide as a repeated nucleotide. This would have two bases attached to a chain that reads.
- Phosphate–sugar–phosphate–sugar: A phosphodiester linkage. Note that these NAD derivatives have a sugar-phosphate–phosphate–sugar sequence, a broader interpretation of dinucleotide terminology.
The reactive center in these compounds relates to the pyridine ring in nicotinamide, which is capable of accepting or donating hydride equivalents according to its oxidation state.
- We have seen that, in biochemical reactions, NADH and NADPH may be considered analogs of complex metal hydride reagents.
- Here is our first example, then, of a nucleotide where the base, nicotinamide, is different from those in nucleic acids.
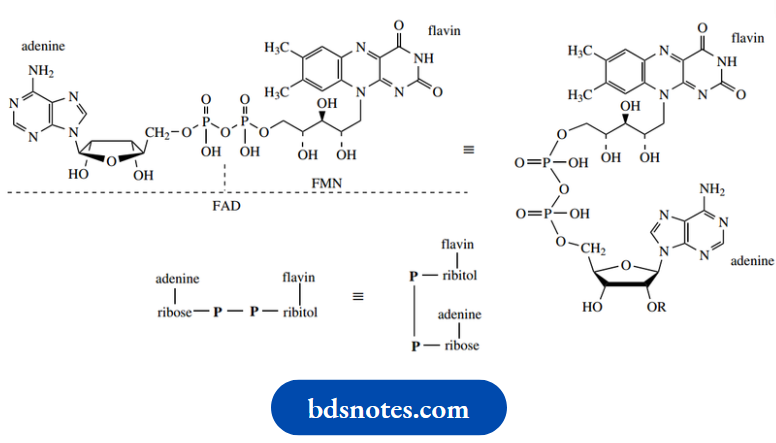
- FAD shares a lot of features with NAD+ and NADP+, but contains two new variants:
- A sugar that is neither ribose nor deoxyribose and a fairly complex heterocyclic base flavin. The new sugar is ribitol, non-cyclic because it contains no carbonyl group.
The chemistry of FAD is concentrated in the flavin part, and features oxidation/reduction processes. FMN, flavin mononucleotide, is simply the flavin-containing
Cyclic AMP
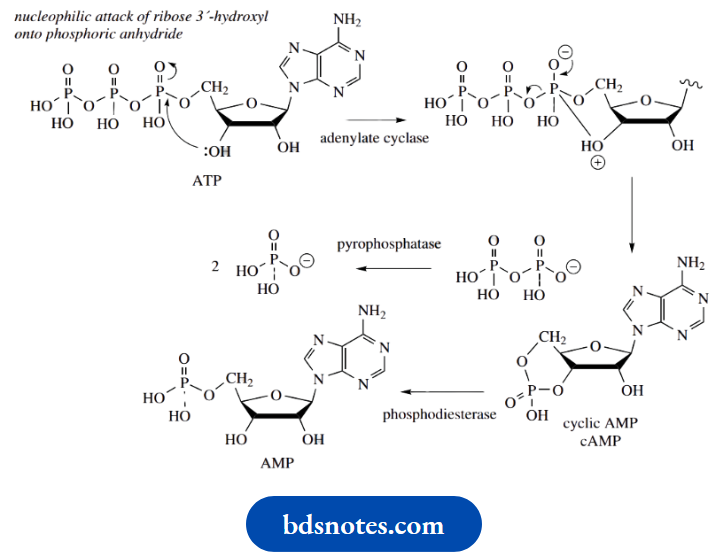
The nucleotide cyclic AMP (3′,5′-cyclic adenosine monophosphate, cAMP) is a cyclic phosphate ester of particular biochemical significance. It is formed from the triester ATP by the action of the enzyme adenylate cyclase, via nucleophilic attack of the ribose 3′-hydroxyl onto the nearest P = O group, displacing diphosphate as the leaving group. It is subsequently inactivated by hydrolysis to 5′-AMP through the action of a phosphodiesterase enzyme
cAMP functions in cells as a second messenger, a mediator molecule that transmits the signal from a hormone. Other second messengers identified include Ca2+, prostaglandins, diacylglycerol, and the equivalent cyclic phosphate derivative of guanosine, cyclic GMP.
- cAMP is the mediator for a variety of drugs, hormones, and neurotransmitters, including adrenaline, glucagon, calcitonin, and vasopressin. Such compounds produce their effects by increasing or decreasing the catalytic activity of adenylate cyclase, thus raising or lowering the cAMP concentration in a cell.
- A pyrophosphatase activity rapidly removes the other reaction product, disturbing the equilibrium, and making the reaction unidirectional cAMP, in turn, is responsible for the activation of various protein kinases that regulate the activity of cellular proteins by phosphorylation of serine and threonine residues using ATP.
The phosphorylated and nonphosphorylated forms of the enzymes catalyze the same reaction, but at quite different rates; often, one of the forms is essentially inactive. This means the activity of enzymes may be switched on or off by addition or removal of phosphate groups, and can thus be controlled by hormones.
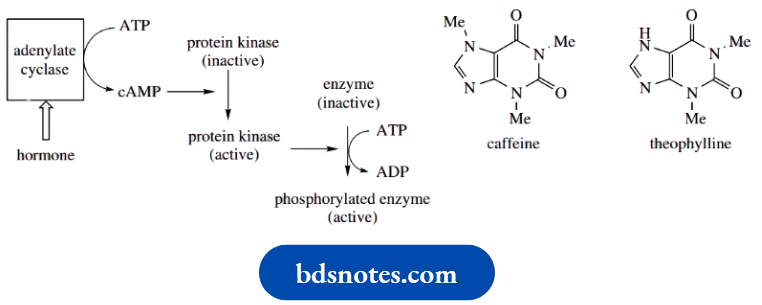
Caffeine in tea and coffee inhibits the phosphodiesterase that degrades cAMP. The resultant increase in cAMP levels, therefore, mimics the action of mediators such as the catecholamines that modulate adenylate cyclase. Caffeine and the related theophylline (both purine alkaloids are thus effective stimulants of the CNS.
Nucleotide Biosynthesis
Nucleic acids are synthesized in nature from nucleoside triphosphates, which are coupled by a chain extension process.
- We have seen that coupling is simply an esterification reaction utilizing the 3′-hydroxyl of the sugar of the growing chain, with diphosphate as a good leaving group.
- Nucleoside triphosphates, especially ATP, have other major biochemical roles. A full discussion of the origins of these compounds is outside our requirements, but there are some features of particular interest pertinent to our understanding of these compounds.
- One of these is that several of the biosynthetic reactions require the involvement of ATP, demonstrating that nucleotide production requires input from other nucleotides.
- Another interesting aspect is the quite different approach nature adopts for the synthesis of pyrimidine or purine nucleotides.
- Pyrimidine nucleotides are made by adding a preformed pyrimidine ring to the sugar-phosphate.
On the other hand, the purine ring of purine nucleotides is built up gradually, and assembly occurs with the growing ring attached to the sugar-phosphate. A common intermediate for all the nucleotides is 5-phosphoribosyl-1-diphosphate (PRPP), produced by successive ATP-dependent phosphorylations of ribose. This has an α-diphosphate leaving group that can be displaced in SN2 reactions.
Similar SN2 reactions have been seen in glycoside synthesis and biosynthesis and for the synthesis of aminosugars. For pyrimidine nucleotide biosynthesis, the nucleophile is the 1-nitrogen of uracil- 6-carboxylic acid, usually called orotic acid. The product is the nucleotide orotidylic acid, which is subsequently decarboxylated to the now-recognizable uridylic acid (UMP).
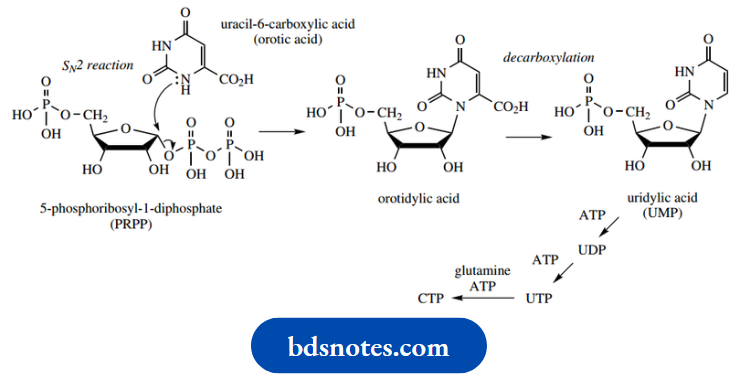
Formation of UTP requires successive phosphorylations using ATP. CTP is, in turn, formed from UTP by an amination reaction in the pyrimidine ring, with the amino acid glutamine supplying the nitrogen; this is also an ATP-dependent reaction.
Glutamine also supplies an amino function to start purine nucleotide biosynthesis. This complex little reaction is again an SN2 reaction on PRPP, but only an amino group from the amide of glutamine is transferred. The product of the enzymic reaction is thus 5-phosphoribosylamine.
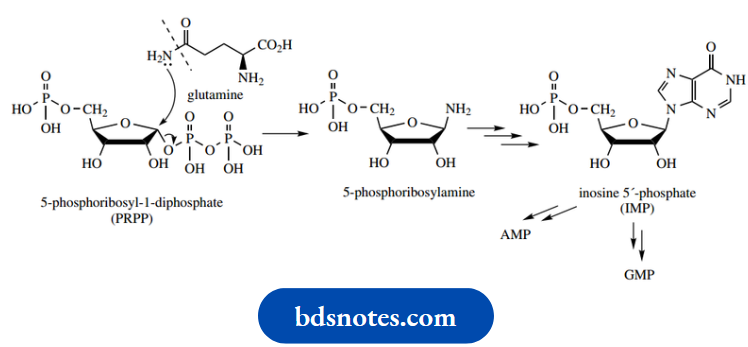
The amino group now provides the nucleus for purine ring formation, an extended series of reactions we shall not describe.
- The first-formed purine product is inosine 5′-phosphate (IMP), which leads to either AMP or GMP; these require amination at alternative sites and utilize either GTP- or ATP-dependent reactions for amination.
- GTP or ATP (as appropriate) will also be required for further phosphorylations to produce the nucleotide triphosphates.
- 2 ‘Deoxyribonucleotides are generally formed by reduction of ribonucleoside diphosphates
- . This involves a series of redox reactions in which NADP+ and FAD play a role, with a subsequent electron transport chain. DNA contains thymine rather than uracil, so thymidine triphosphate (dTTP) is a requirement.
- Methylation of dUMP to dTMP is a major route to thymine nucleotides and is dependent upon N5, N10-methylenetetrahydrofolate as the source of the methyl group.
Determination Of Nucleotide Sequence
Restrictions on endonucleases
Natural DNA molecules are extremely large, and for sequence determination, it is necessary to cleave them into manageable fragments. This may be accomplished by using enzymes, called restriction endonucleases, which are obtained mainly from bacterial sources.
- These enzymes appear to have developed so that a cell can destroy foreign, particularly viral, DNA.
- The enzymes, of which several hundred are available, cleave the DNA at specific points in the chain, dictated by a series of nucleotides, typically three to six nucleotides.
- For example, the enzyme EcoRI (from Escherichia coli ) cleaves a GAATTC sequence between G and A.
- Also important is the property that most restriction endonucleases cleave both strands of DNA because the recognition sequence reads the same both ways: the complementary strand to GAATTC (in 5’→ 3’direction) is CTTAAG (in 3’→ 5’direction).
- The restriction endonuclease recognizes a specific sequence, but the probability of these sequences occurring in a given DNA molecule is usually quite low; therefore, cleavage produces only a few fragments.
- The use of a different enzyme on the same DNA will produce different fragments, but there then will be an overlap of sequences.
- Hence, sequencing of both sets of fragments should allow the full sequence to be deduced. This deductive approach is thus similar to that used in amino acid sequencing.
Chemical sequencing
Before separation, double-stranded restriction fragments are labeled chemically, by attaching a radioactive or fluorescent marker to the 5′-end of the chain.
For example: Radioactive 32P-labelled phosphate may be added using labeled ATP in an enzymic reaction.
- The labeled fragments are then separated chromatographically using conditions that are known to cause strand separation into single-stranded DNA molecules.
- The separated fragments are then split into four portions, and each portion is treated chemically with a suitable reagent.
- The reagent needs to induce cleavage reactions, but it shows selectivity for the different nucleotides.
- Now this could potentially lead to almost total cleavage, but the trick is to use reagents at concentrations so low that, statistically, only one cleavage occurs per chain. The reagents are dimethyl sulfate and hydrazine (only two reagents, but read on), and though we shall not consider the full mechanisms of the reactions here.
They may be summarized as follows
- Me2SO4, then aqueous piperidine; cleavage at G;
- Me2SO4 and aqueous formic acid, then aqueous piperidine; cleavage at A and G;
- Aqueous hydrazine (H2NNH2), then aqueous piperidine; cleavage at C and T;
- Aqueous hydrazine (H2NNH2) and NaCl, then aqueous piperidine; cleavage at C
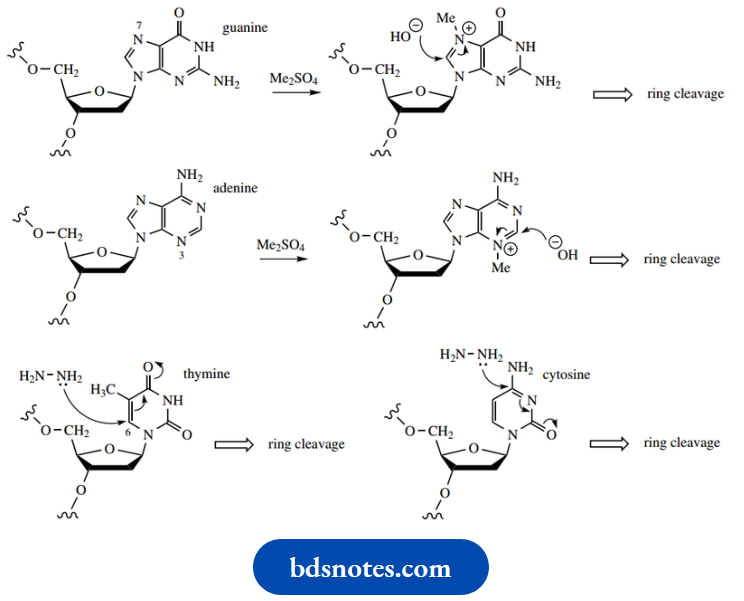
Dimethyl sulfate
Dimethyl sulfate is an effective methylating agent Methylation of the purine rings in guanine and adenine makes them susceptible to hydrolysis and subsequent rupture. This, in turn, makes the glycosidic bond vulnerable to attack, and the heterocycle is displaced from the phosphodiester. The phosphodiester bond can then be cleaved by basic hydrolysis (aqueous piperidine).
- Guanine is methylated on the imidazole ring at N-7, whereas adenine undergoes methylation at N-3.
- Under the conditions used, guanine is methylated more readily than adenine; therefore, cleavage of the DNA occurs predominantly where there a guanine residues.
- However, by treating the methylated DNA with acid, cleavage at the methylated adenine sites becomes enhanced, and the chain is broken at sites that originally contained either adenine or guanine.
- The pyrimidines cytosine and thymine both react with hydrazine, which initially attacks the unsaturated carbonyl system and then leads to ring opening.
Again, base treatment is used to hydrolyze the phosphodiester bond.
This reaction becomes selective for cytosine in the presence of NaCl, which suppresses the reaction with thymine.
- The reaction products from the four reactions are then separated by gel electrophoresis in parallel lanes.
- This procedure will separate the components according to their charge (mainly from phosphate groups) and their size.
- The smallest species will migrate furthest. After chromatography, the gel is visualized by autoradiography, detecting bands via the radioactive tracer used. The base sequence can be read directly from the gel by the pattern of bands produced using the following reasoning
Consider a short sequence as shown (by convention written from 5′-end to 3′-end):
AGTCGGAACGTA
This is labelled at the 5’end with 32P to give
32P–AGTCGGAACGTA
Cleavage at the 5′-side of G residues using the first reagent (Me2SO4, then aqueous piperidine) leads to fragments
32P – A
32P – AGTC
32
P – AGTCG
32P – AGTCGGAAC
Of course, there will be other fragments that do not contain the 5′-end with its 32P label, but we shall not detect any of these since they contain no radioactive label.
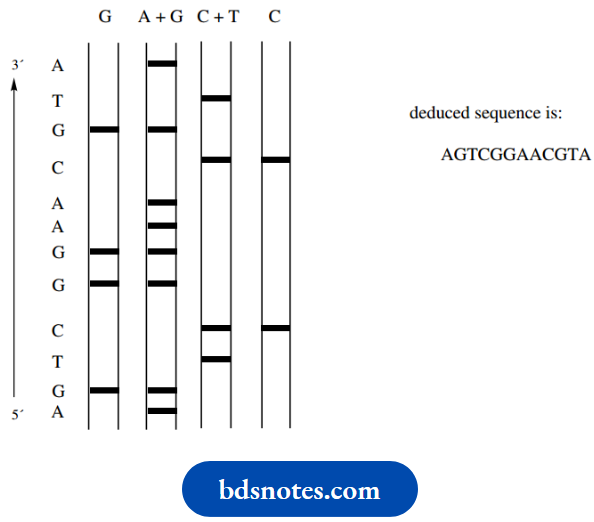
Corresponding fragments will be produced when we use the other three types of cleavage reactions. The resultant chromatogram with the four reaction mixtures will then look something like though the bands will be much closer together in practice.
Bands that occur in the left-hand lane represent guanine, and bands that occur in the second lane but not the first lane represent adenine. Similarly, bands in the third lane but not the fourth lane represent thymine, and bands that occur in the fourth lane represent cytosine. By reading up the chromatogram, the sequence AGTCGGAAC may be deduced. It is possible to distinguish about 200 bands on a single gel.
The process is so reliable that automated equipment is available to perform routine analyses. As an alternative to using radioactive labeling, a modification uses differently colored fluorescent dyes, one for each base-selective reaction. All samples are then applied in one lane, and the base sequence can then be read automatically from the color of the bands along the gel. Similar sequencing methodology can be applied to RNA samples.
Oligonucleotide Synthesis: The phosphoramidite Method
The ability to synthesize chemically short sequences of single-stranded DNA (oligonucleotides) is an essential part of many aspects of genetic engineering.
The method most frequently employed is that of solid-phase synthesis, where the basic philosophy is the same as that in solid-phase peptide synthesis.
- In other words, the growing nucleic acid is attached to a suitable solid support, protected nucleotides are supplied in the appropriate sequence, and each addition is followed by repeated coupling and deprotection cycles.
- As with peptide synthesis, similar considerations must be incorporated into the methodology. Vulnerable functional groups in the base, the sugar, and the phosphates will need to be protected.
- The groups to be coupled may need suitable activation, and after the coupling reaction, the protecting groups must be removed under mild conditions.
- In addition, we need to attach the starting material to the support, and eventually, the product will need to be released from the support.
Nevertheless, the procedure is efficient and has allowed the development of automatic DNA synthesizers capable of preparing oligonucleotides of up to about 150 residues.
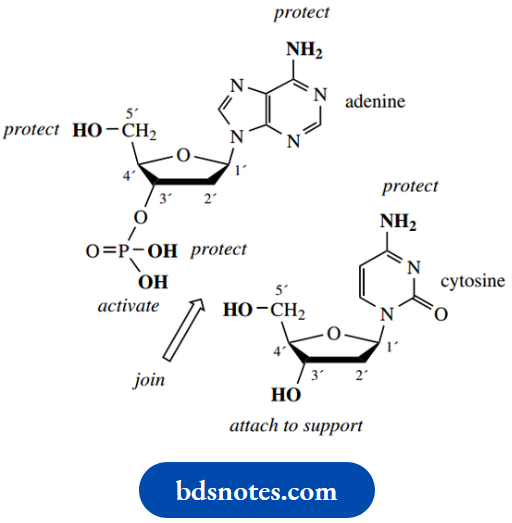
In solid-phase syntheses, oligonucleotides are usually synthesized in the 5′-direction from an immobilized 3′-terminus
- The solid phase is generally silica or controlled pore glass (CPG), which has been derivatized to provide a spacer molecule carrying a primary amino group. This spacer group is used to bring the nucleotide away from the support and allow the reagents free access.
- The first residue, as a nucleoside (i.e. without phosphate), is affixed to the support via its 3′-hydroxyl, using a succinic acid residue to achieve bonding, and also extend the spacer further.
The succinic acid residue thus has an amide link at one end and an ester link to the sugar of the nucleoside. In practice, the ester linkage is performed first.
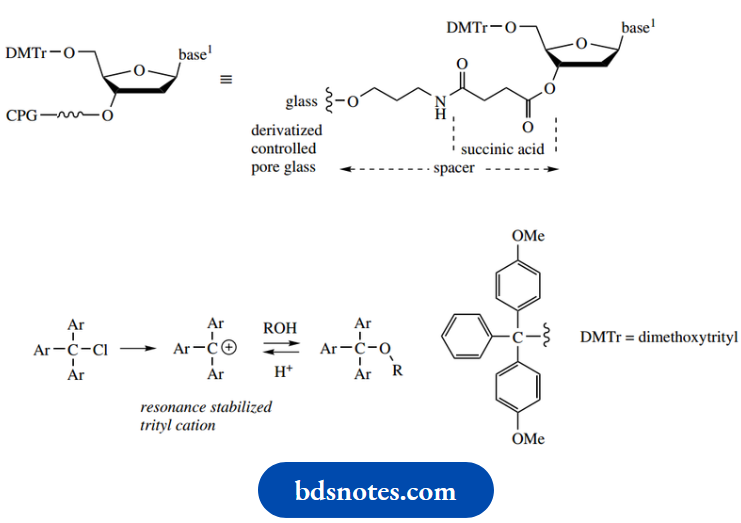
Protection
Protection of the 5′-hydroxyl of the sugar unit is usually as a dimethoxytrityl ether (trityl: triphenyl methyl), by reaction with dimethoxytrityl chloride
. The dimethoxytrityl group is bulky, and the reaction only occurs at the primary 5′-hydroxyl of the sugar group, the secondary 3′-hydroxyl being too hindered to react.
- This protecting group is easily removed by treatment with acid, even more easily than trityl groups, since the electron-donating methoxy groups stabilize the triaryl methyl carbocation that is an intermediate in the deprotection reaction
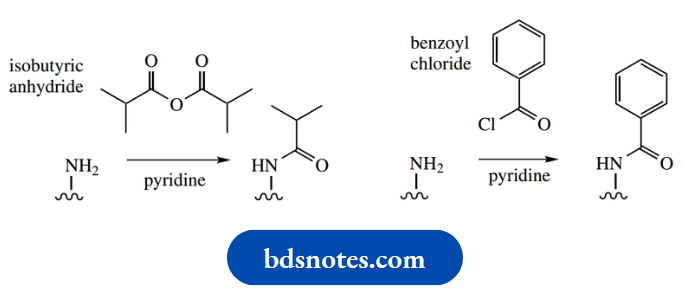
- The bases adenine, guanine, and cytosine all contain exocyclic amino substituents that require protection since these are potential nucleophiles
- . They are converted into amides that are stable to the other reagents used in the process, yet can be removed readily by basic hydrolysis.
The most effective protecting groups are isobutyryl for the amino group of guanine and benzoyl for adenine and cytosine. Thymine has no exocyclic nitrogen and does not need protection.
Protection and activation of the phosphate moiety is achieved by employing a phosphoramidite derivative, –P(OR)NR2. This reagent has phosphorus in its PIII oxidation state; the phosphate that we finally require contains PV. Favoured R groups in the phosphoramidite are 2-cyanoethyl for OR and 2-propyl (isopropyl) for NR2. The reagent used to attach this to the 3′-hydroxyl is the phosphorodiamidate shown, the hydroxyl displacing an NR2 group in the presence of tetrazole as a mild acidic catalyst.
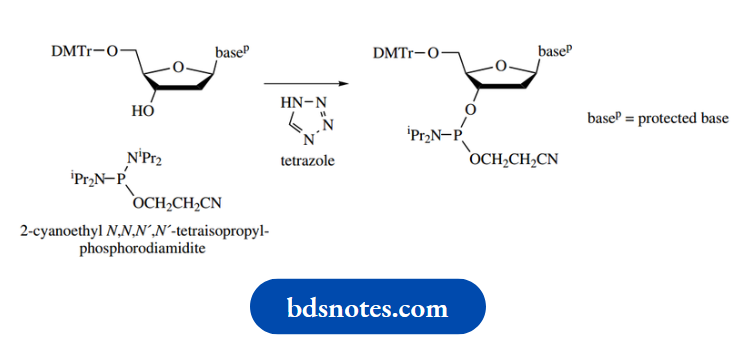
In what is essentially a repeat of this reaction, the 5’hydroxyl of a second nucleoside can couple to this intermediate; this is the crucial coupling reaction in the sequence shown below.
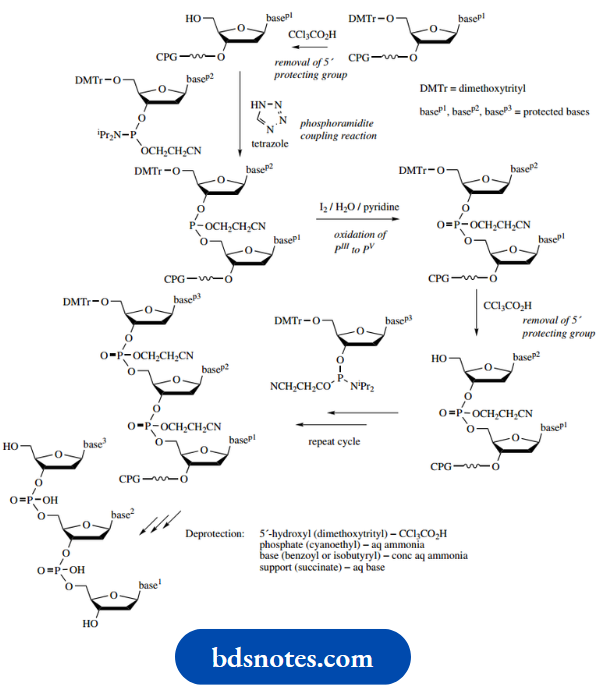
Of course, the product does not have a phosphate linker between the two nucleosides, and phosphorus is still in the wrong oxidation state.
- This is remedied by oxidation of the dinucleotide phosphite to a phosphotriester using iodine. We now have the required phosphate linker, though it is still protected with the cyanoethyl group.
- This is retained at this stage. The dimethoxytrityl ester-protecting group is now removed by treatment with a mild acid (CCl3CO2H), which is insufficiently reactive to hydrolyze the amide protection of bases or the cyanoethyl protection of the phosphate.
- The coupling cycle can now be repeated using a phosphoramidite derivative of the next appropriate nucleoside. The sequences will be continued as necessary until the desired oligonucleotide is obtained.
- It then remains to remove protecting groups and release the product from the support.
- All of these tasks, except for the removal of the dimethoxytrityl group, are achieved by the use of a single deprotection reagent, an aqueous base (ammonia).
- The cyanoethyl groups are lost from the phosphates by base-catalyzed elimination, and amide protection of the bases is removed by base-catalyzed hydrolysis. The latter process also achieves hydrolysis of the succinate ester link to the support.
Copying DNA: The Polymerase Chain Reaction
The polymerase chain reaction (PCR), developed by Mullis, is a simple and most effective way of amplifying, i.e. producing multiple copies of, a DNA sequence. It finds applications in all sorts of areas not immediately associated with nucleic acid biochemistry,
Examples:
Genetic screening, medical diagnostics, forensic science, and evolutionary biology.
The general public is now well aware of the importance of some of these topics.
Example:
The ability to identify a person by DNA analysis, but perhaps does not realize that tiny samples of DNA must be copied millions of times to provide a sample large enough for chromatographic analysis.
- PCR makes use of the heat-stable enzyme DNA polymerase from the bacterium Thermus aquaticus and its ability to synthesize complementary strands of DNA when supplied with the necessary deoxyribonucleoside triphosphates. We have already looked at the chemistry of DNA replication, and this process is the same, though it is carried out in the laboratory and has been automated.
- Although knowledge of the whole nucleotide sequence of the target area of DNA is not required, one must know the sequence of some small stretch on either side of the target area. These data may be known from other sequencing studies; or, surprisingly, it can even be predictable from knowledge of related genes.
- Two single-stranded oligonucleotides, one for each sequence, are then synthesized to act as primers.
- Typically, the primers should contain about 20 nucleotides, and they must be complementary to the DNA sequences of opposite strands.
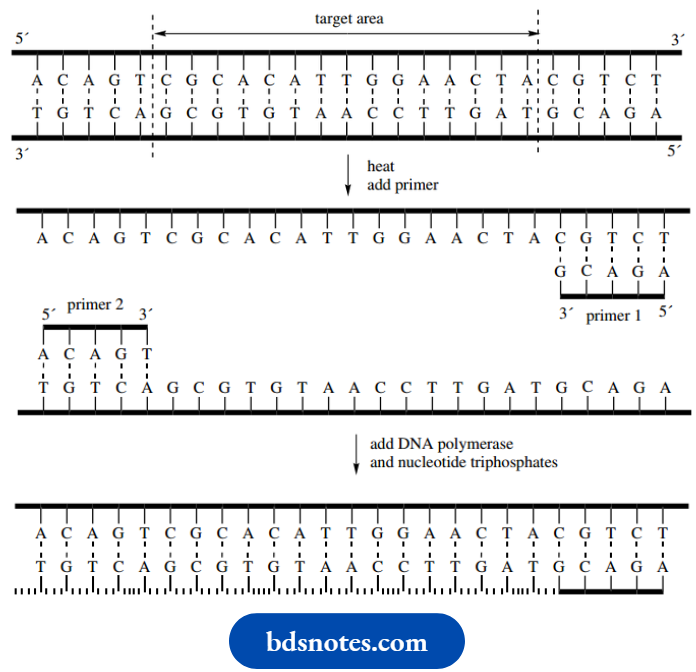
- In the schematic illustration of the process, the central target area is indicated, and the primers are depicted as short complementary sequences.
Initially, the double-stranded DNA is heated to separate the strands. The primers are then added and the temperature is lowered so that the primers anneal to the complementary sequences of each strand. In the presence of nucleoside triphosphates, the DNA polymerase enzyme will replicate a length of DNA starting from the 3′-end of a nucleotide, extending the chain towards the 5′-end.
It will thus start chain extension from the 3′-ends of the primers and continue to the end of the DNA strands. This will lead to two double-stranded DNA molecules, composed of initial strands, and primer plus newly synthesized DNA, as shown.
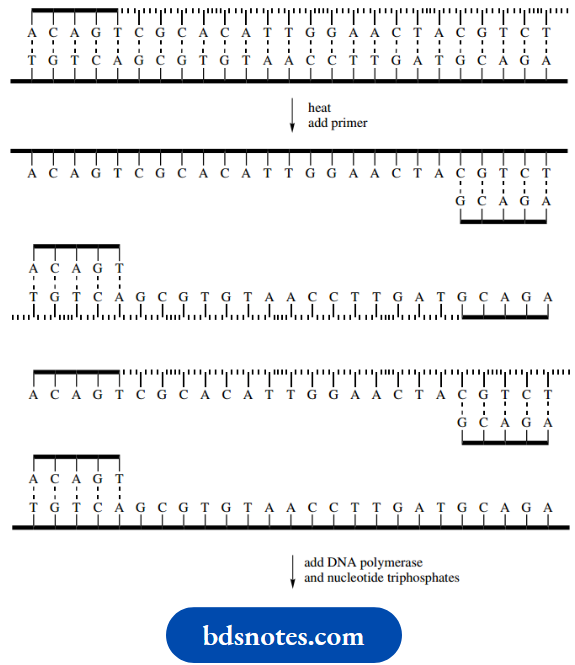
The process is repeated. Heating causes the separation of strands, and cooling allows the primer to attach to the appropriate nucleotide sequence.
- Enzymic chain extension then produces four double-stranded DNA molecules.
- The number of DNA molecules doubles in each cycle of the process, so that after 30 cycles, say, we have 230 molecules (approximately 109 copies).
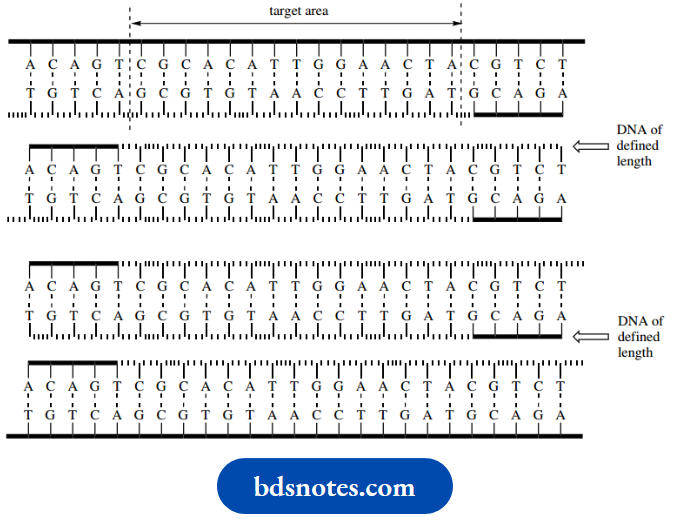
- However, there is another, less obvious feature that makes the PCR even more useful. In the second cycle, two of the newly synthesized single-stranded chains will be of defined length.
- They will consist of the target area plus two primers; the 5’ends of the primers define the length of DNA.
- Other molecules will be much longer because replication goes on to the end of the template.
- Should you wish to follow this through, you will find that, after the third cycle, there will be eight single-stranded DNA molecules of defined length and eight that are longer.
- After each cycle, the number of defined-length DNA molecules increases geometrically, whereas the number of DNA strands containing sequences outside of the primers only increases arithmetically. This means that, after about 20 cycles, the DNA synthesized is almost entirely composed of molecules whose length is defined by the primers, i.e. the target area plus a short extra length defined by the primers
Leave a Reply